Md Simulations Modified Pdf

Md Simulations Modified Pdf The purpose of this project is to perform md simulations for short peptides and explore their secondary structure propensities. two peptides, acetyl vvvv nh2 and acetyl aaaaaaaaa nh2, are to be used in the simulations. This review synthesizes principles underlying md simulations, computational methodologies, and their applications across biomolecular research and materials science.
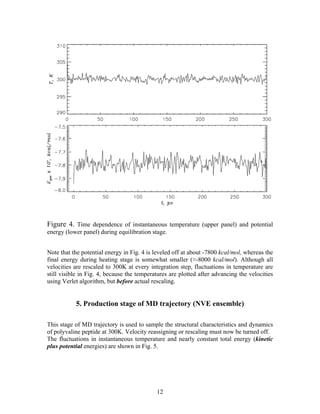
Md Simulations Modified Pdf The document describes the steps to set up and run a 1 ns molecular dynamics simulation of a polyvaline peptide using namd. We used simulations to determine where this molecule binds to its receptor, and how it changes the binding strength of molecules that bind elsewhere (in part by changing the protein’s structure). we then used that information to modify the molecule such that it has a different effect. This review synthesizes principles underlying md simulations, computational methodologies, and their applications across biomolecular research and materials science. This review synthesizes principles underlying md simulations, computational methodologies, and their applications across biomolecular research and materials science, and highlights recent advancements and explores future directions.

Md Simulations Modified Pdf This review synthesizes principles underlying md simulations, computational methodologies, and their applications across biomolecular research and materials science. This review synthesizes principles underlying md simulations, computational methodologies, and their applications across biomolecular research and materials science, and highlights recent advancements and explores future directions. The best way is to made md simulation and gather needed data in file, then we have all required data to be used in calculation of time correlation. Md simulation mimics the changes in the structures of biological molecules over a given period of time, giving us atomic insights about the change in structure. Molecular dynamics (md) simulation. • computer simulation method which simulates dynamic evolution of a microscopic system (ie. physical movements of atoms and molecules), . the behavior and properties of the system are then analyzed. typical size of system can be investigated: <10. 6. atoms. 2. We used simulations to determine where this molecule binds to its receptor, and how it changes the binding strength of molecules that bind elsewhere (in part by changing the protein’s structure).

Model Used In The Md Simulations Download Scientific Diagram The best way is to made md simulation and gather needed data in file, then we have all required data to be used in calculation of time correlation. Md simulation mimics the changes in the structures of biological molecules over a given period of time, giving us atomic insights about the change in structure. Molecular dynamics (md) simulation. • computer simulation method which simulates dynamic evolution of a microscopic system (ie. physical movements of atoms and molecules), . the behavior and properties of the system are then analyzed. typical size of system can be investigated: <10. 6. atoms. 2. We used simulations to determine where this molecule binds to its receptor, and how it changes the binding strength of molecules that bind elsewhere (in part by changing the protein’s structure).
Comments are closed.