Molecular Dynamics Simulation Explained

Molecular Dynamics Simulation Accuracy Speed Predictive Power Molecular dynamics (md) is a computer simulation method for analyzing the physical movements of atoms and molecules. the atoms and molecules are allowed to interact for a fixed period of time, giving a view of the dynamic "evolution" of the system. Molecular dynamics is defined as a simulation technique that generates the dynamic evolution of atoms in a molecular system over time by calculating atomic positions, velocities, and accelerations using newton's laws of motion.
Getting Started With Molecular Dynamics Simulation Insilicosci We used simulations to determine where this molecule binds to its receptor, and how it changes the binding strength of molecules that bind elsewhere (in part by changing the protein’s structure). Learn the basics of molecular dynamics simulations, a powerful technique used to study atoms and molecules in different systems and their applications. This article explains how atomic motion is tracked by numerically solving equations of motion, the interatomic potential that is key to the simulation, and noteworthy machine learning force fields. you can learn about the fundamentals, applications, and latest technologies of molecular dynamics. Explain the theory and mathematics behind a molecular dynamics simulation, including approximations and sources of inaccuracies. describe what a force field is and what its parameters are.

Molecular Dynamics Simulation Protac This article explains how atomic motion is tracked by numerically solving equations of motion, the interatomic potential that is key to the simulation, and noteworthy machine learning force fields. you can learn about the fundamentals, applications, and latest technologies of molecular dynamics. Explain the theory and mathematics behind a molecular dynamics simulation, including approximations and sources of inaccuracies. describe what a force field is and what its parameters are. In essence, md is a computational method that models how atoms and molecules move over time by applying classical newtonian physics to a detailed molecular model. think of it like creating a. The impact of molecular dynamics (md) simulations in molecular biology and drug discovery has expanded dramatically in recent years. these simulations capture the behavior of proteins and other biomolecules in full atomic detail and at very fine temporal resolution. In simulations the “true” pair potential is usually replaced by an effective pair potential, which incorporate the average many body e ects into the first term :. Molecular dynamics (md) is a computational technique used to analyse the physical behaviour of material systems at the atomic scale. in an md approach, the material system is considered as a group of rigid bodies (atoms), wherein the dynamic evolution of atoms is.

Molecular Dynamics Simulation Model Download Scientific Diagram In essence, md is a computational method that models how atoms and molecules move over time by applying classical newtonian physics to a detailed molecular model. think of it like creating a. The impact of molecular dynamics (md) simulations in molecular biology and drug discovery has expanded dramatically in recent years. these simulations capture the behavior of proteins and other biomolecules in full atomic detail and at very fine temporal resolution. In simulations the “true” pair potential is usually replaced by an effective pair potential, which incorporate the average many body e ects into the first term :. Molecular dynamics (md) is a computational technique used to analyse the physical behaviour of material systems at the atomic scale. in an md approach, the material system is considered as a group of rigid bodies (atoms), wherein the dynamic evolution of atoms is.

Molecular Dynamics Simulation Model Download Scientific Diagram In simulations the “true” pair potential is usually replaced by an effective pair potential, which incorporate the average many body e ects into the first term :. Molecular dynamics (md) is a computational technique used to analyse the physical behaviour of material systems at the atomic scale. in an md approach, the material system is considered as a group of rigid bodies (atoms), wherein the dynamic evolution of atoms is.
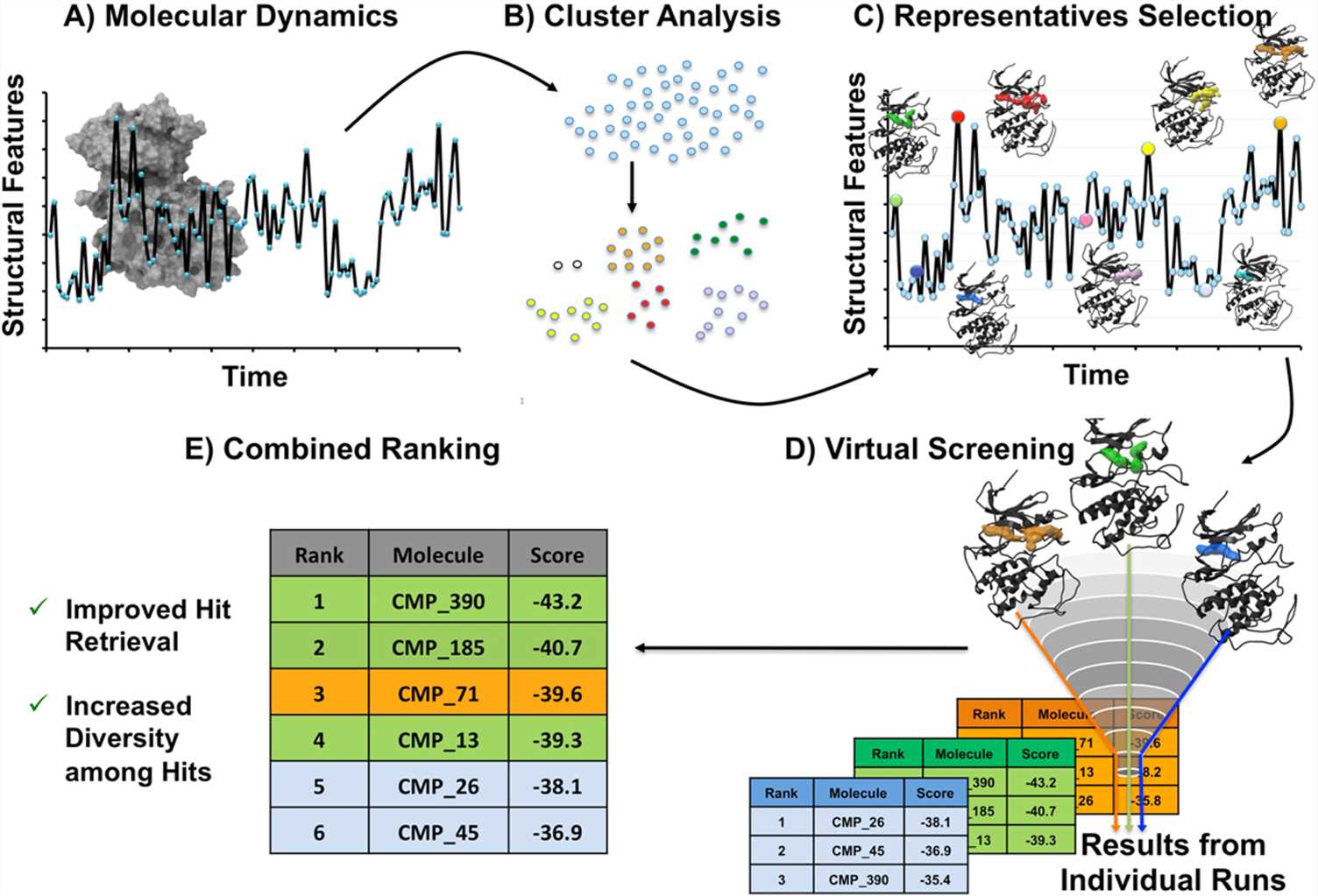
Molecular Dynamics Simulation Creative Biostucture Drug Discovery
Comments are closed.