Genomicranges Seqinfo V1
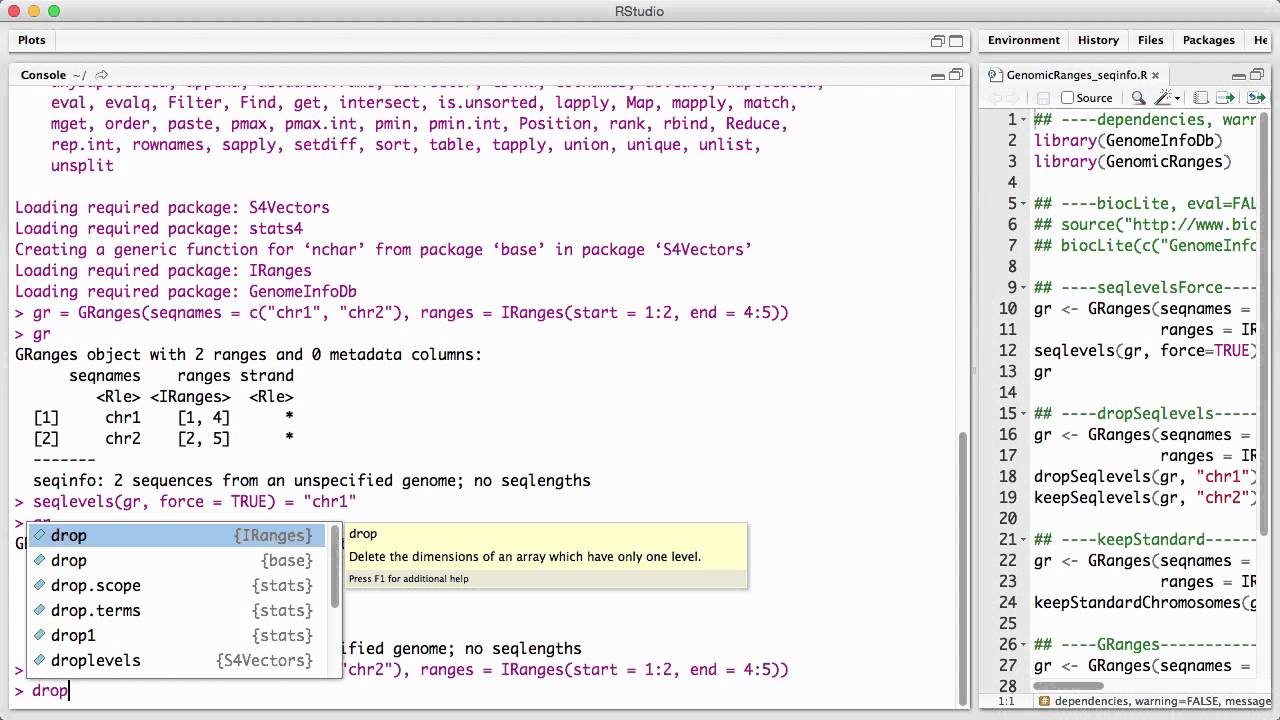
Genomicranges Seqinfo V1 Youtube The genomicranges package defines general purpose containers for storing and manipulating genomic intervals and variables defined along a genome. The granges class contains seqinfo information about the length and the names of the chromosomes. here we will briefly discuss strategies for harmonizing this information.
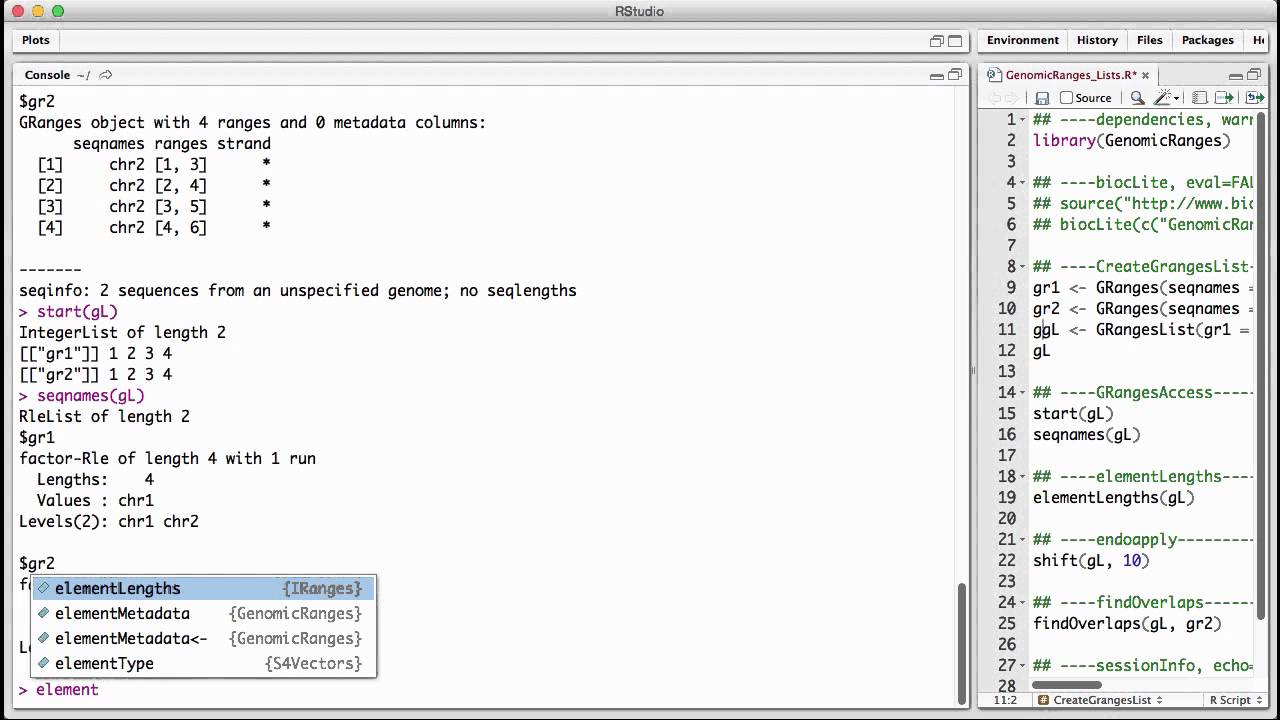
Genomicranges Lists V1 Youtube Genomicranges is an r bioconductor package for representing and manipulating genomic intervals. see bioconductor.org packages genomicranges for more information including how to install the release version of the package (please refrain from installing directly from github). Genomicranges: representation and manipulation of genomic intervals. the ability to efficiently represent and manipulate genomic annotations and alignments is playing a central role when it comes to analyzing high throughput sequencing data (a.k.a. ngs data). The genomicranges package defines seqinfo and seqinfo< methods for these low level data types: list, rangeslist and rangeddata. those objects do not have the means to formally store sequence information. thus, the wrappers simply store the seqinfo object within metadata(x). Bioconductor for genomic data science: kasperdanielhansen.github.io genbioconductor.
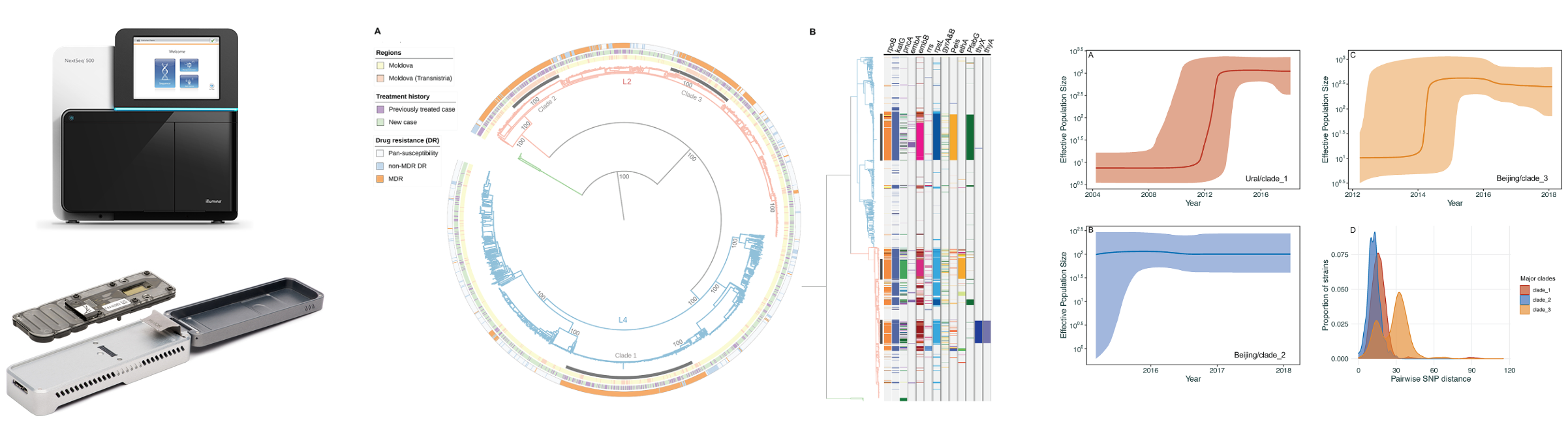
Home Genomics Course The genomicranges package defines seqinfo and seqinfo< methods for these low level data types: list, rangeslist and rangeddata. those objects do not have the means to formally store sequence information. thus, the wrappers simply store the seqinfo object within metadata(x). Bioconductor for genomic data science: kasperdanielhansen.github.io genbioconductor. [docs] defset seqnames(self,seqnames:sequence[str],in place:bool=false) >"seqinfo":""" args: seqnames: list of sequence names, of length equal to the number of names in this ``seqinfo`` object. all names should be unique strings. in place: whether to modify the ``seqinfo`` object in place. In most cases, you don't need to download the package archive at all. the ability to efficiently represent and manipulate genomic annotations and alignments is playing a central role when it comes to analyze high throughput sequencing data (a.k.a. ngs data). the package defines general purpose containers for storing genomic intervals. This man page documents list ("inter range transformations") of a genomicranges object (i.e. of an object that belongs to the genomicranges class or one of its subclasses, this includes for example granges objects), or a grangeslist object. The seqinfo class stores the names, lengths, circularity flags, and genomes for a particular collection of sequences. these sequences are typically the chromosomes and or scaffolds of a specific genome assembly of a given organism.
Comments are closed.