Genomicranges Granges Usage V1

Tidy Genomicranges From Michael Love In this lecture note, we demonstrate how to specify regions of the genome (genomic ranges) in r bioconductor. we will make use of the genomicranges package and the granges object defined by this package. the package also provides a number of methods that work on this type of object. Think of this as an expanded and more versatile class. ## granges, metadata `granges` (unlike `iranges`) may have associated metadata. this is immensely useful.
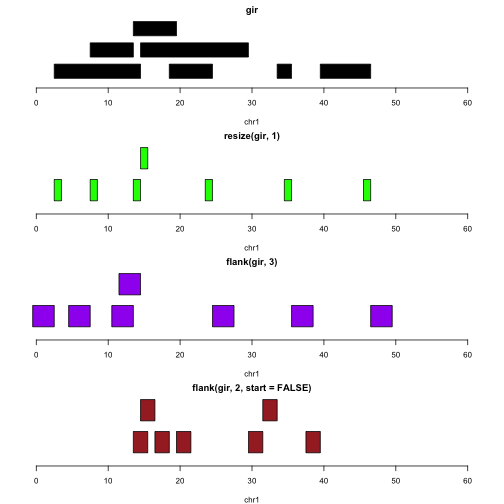
Iranges And Granges In this article, we'll explore what genomicranges offers how to use it effectively, and examples of its application. what is genomicranges? the genomicranges is a bioconductor package tailored for the handling of genomic intervals and sequences. Granges: single interval range features splitting and combining granges objects subsetting granges objects basic interval operations for granges objects. Value an iranges object for absoluteranges. a granges object for relativeranges. absoluteranges and relativeranges both return an object that is parallel to the input object (i.e. same length and names). Here are some simple usecases with pseudo code showing how we can accomplish various tasks with granges objects and functionality. suppose we want to identify transcription factor (tf) binding sites that overlaps known snps. pseudocode: (watch out for strand).

An Illustration Of The Granges Data Model For A Sample From An Rna Seq Value an iranges object for absoluteranges. a granges object for relativeranges. absoluteranges and relativeranges both return an object that is parallel to the input object (i.e. same length and names). Here are some simple usecases with pseudo code showing how we can accomplish various tasks with granges objects and functionality. suppose we want to identify transcription factor (tf) binding sites that overlaps known snps. pseudocode: (watch out for strand). Two of the most common data structures for this purpose are iranges and granges, available from the iranges and genomicranges packages, respectively. Bioconductor for genomic data science: kasperdanielhansen.github.io genbioconductor. Description the granges class is a container for the genomic locations and their associated annotations. Granges is a vector of genomic locations and associated annotations. each element in the vector is comprised of a sequence name, an interval, a strand, and optional metadata columns (e.g. score, gc content, etc.).
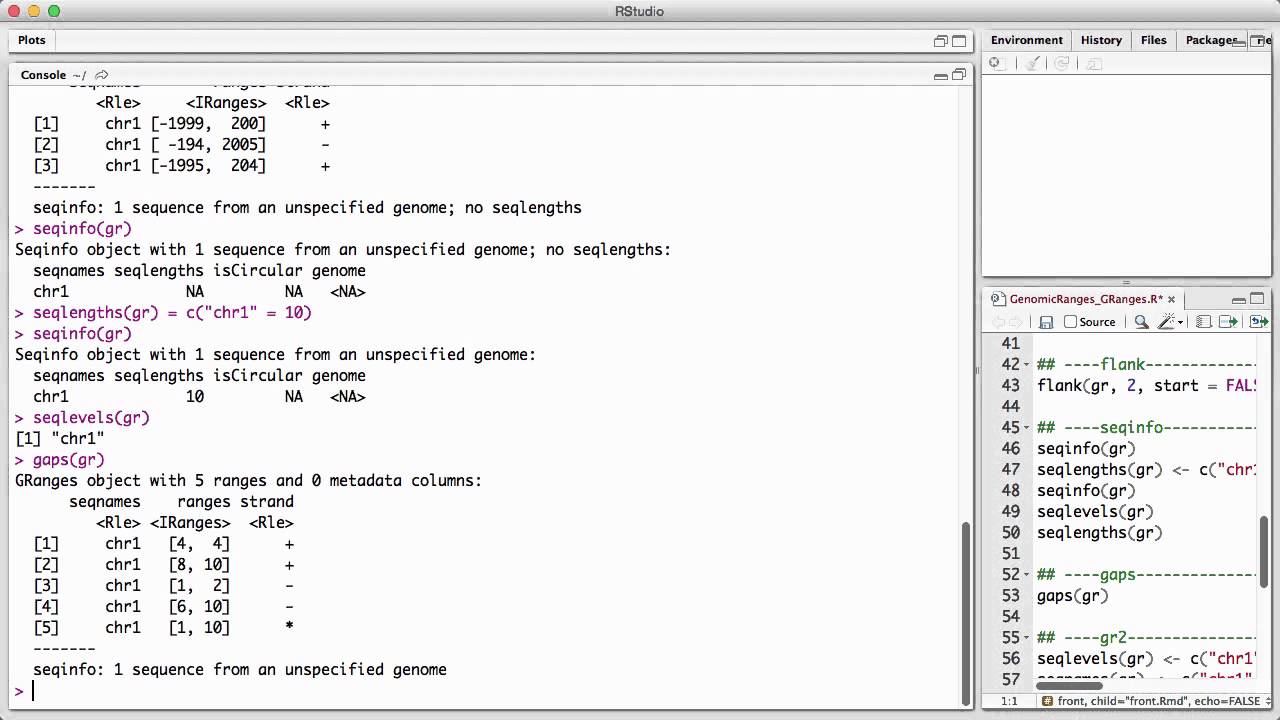
Genomicranges Granges V1 Youtube Two of the most common data structures for this purpose are iranges and granges, available from the iranges and genomicranges packages, respectively. Bioconductor for genomic data science: kasperdanielhansen.github.io genbioconductor. Description the granges class is a container for the genomic locations and their associated annotations. Granges is a vector of genomic locations and associated annotations. each element in the vector is comprised of a sequence name, an interval, a strand, and optional metadata columns (e.g. score, gc content, etc.).
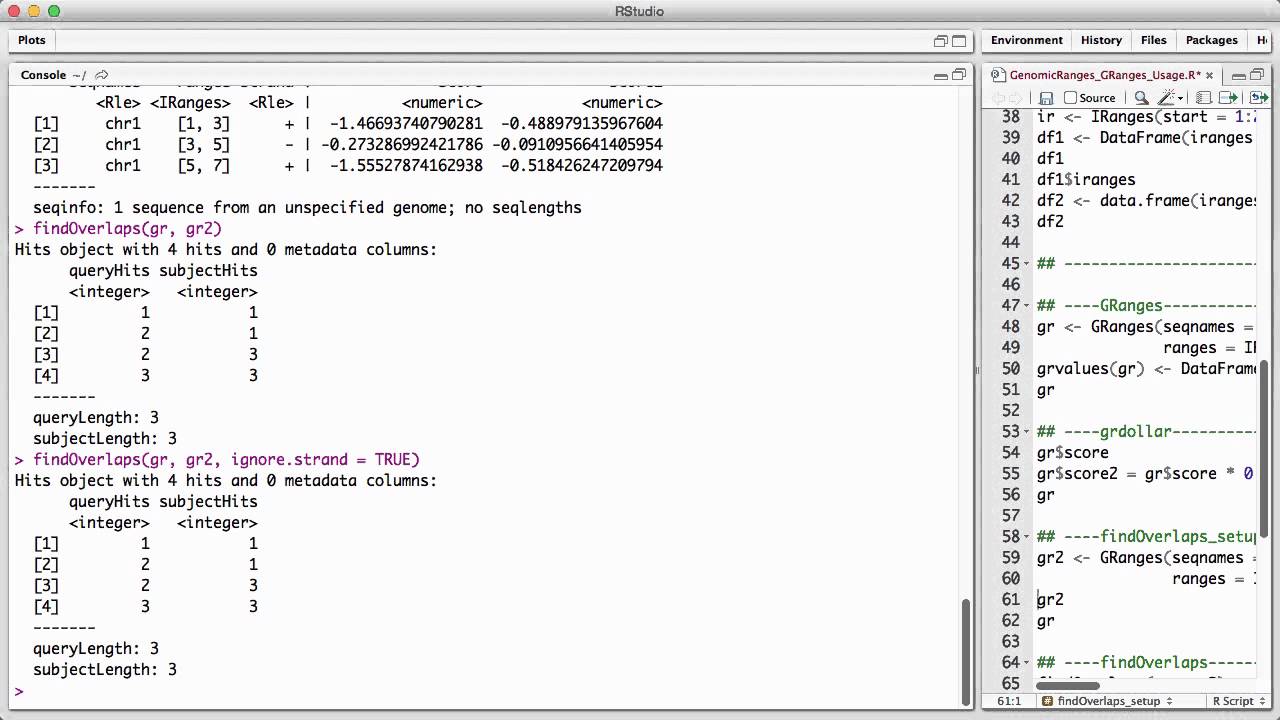
Genomicranges Granges Usage V1 Youtube Description the granges class is a container for the genomic locations and their associated annotations. Granges is a vector of genomic locations and associated annotations. each element in the vector is comprised of a sequence name, an interval, a strand, and optional metadata columns (e.g. score, gc content, etc.).
Comments are closed.