Genomicranges Granges V1 Youtube

Couple Ouvert Live à Trois Rivières Avec Charles Antoine Des Granges Bioconductor for genomic data science: kasperdanielhansen.github.io genbioconductor. In this lecture note, we demonstrate how to specify regions of the genome (genomic ranges) in r bioconductor. we will make use of the genomicranges package and the granges object defined by this package. the package also provides a number of methods that work on this type of object.

Session Granges Youtube In this article, we'll explore what genomicranges offers how to use it effectively, and examples of its application. what is genomicranges? the genomicranges is a bioconductor package tailored for the handling of genomic intervals and sequences. This package lays a foundation for genomic analysis by introducing three classes (granges, gpos, and grangeslist), which are used to represent genomic ranges, genomic positions, and groups of genomic ranges. this vignette focuses on the granges and grangeslist classes and their associated methods. On this level we will learn iranges and granges which simply record start and stop positions of features such as atacseq peaks where they start and stop in. Granges (from genomicranges package) is the main object that holds the genomic intervals and extra information about those intervals. here we will show how to create one.

Granges Overview V1 Youtube On this level we will learn iranges and granges which simply record start and stop positions of features such as atacseq peaks where they start and stop in. Granges (from genomicranges package) is the main object that holds the genomic intervals and extra information about those intervals. here we will show how to create one. How to simply change chromosome names of a granges object? how to build a grangeslist where each granges element is a cds coordinate of gene transcripts? failed to get promoters using promoters () in r 4.0.3. Access information and perform operations on granges with functions from the genomicranges library. the genomation library has functions to easily load annotation files and other genomics datasets. Bondi gets news she fearedweekend update: chad maxxington on the art of looksmaxxing. Here we are going to move from iranges to granges. the key difference is that granges keeps information in "meta". the key is that the chromosome number is included in the start and stop.
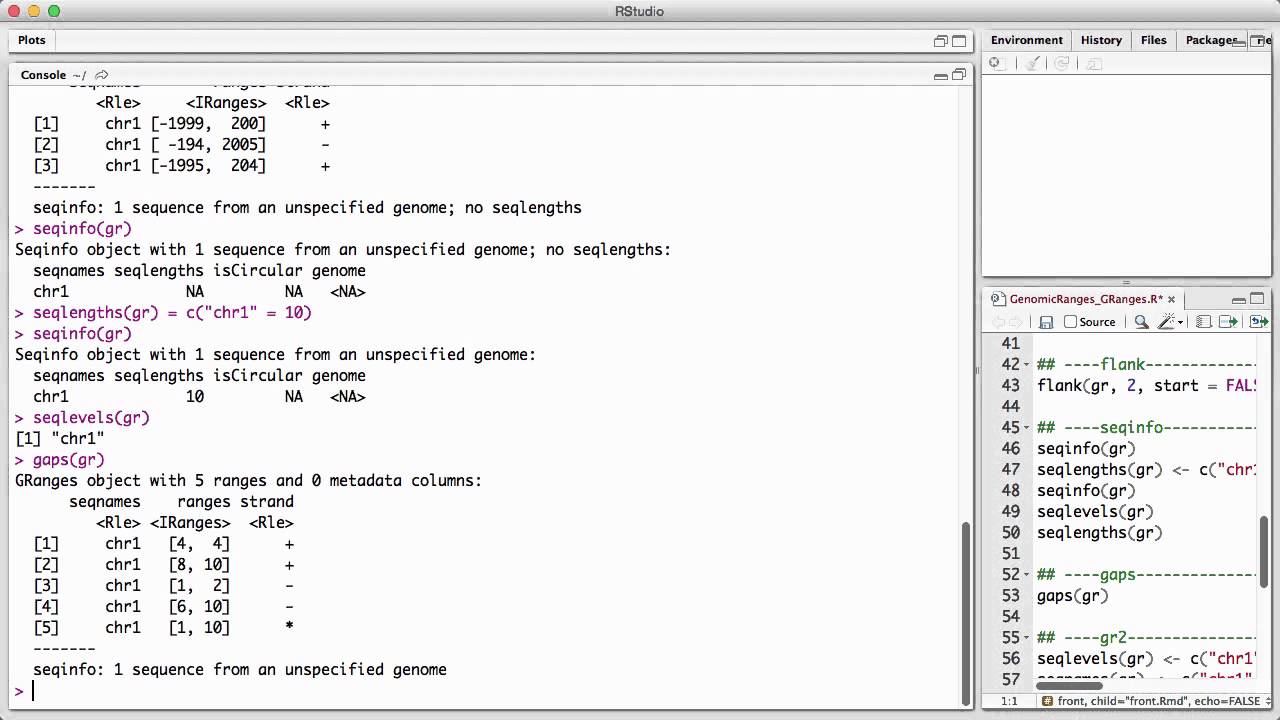
Genomicranges Granges V1 Youtube How to simply change chromosome names of a granges object? how to build a grangeslist where each granges element is a cds coordinate of gene transcripts? failed to get promoters using promoters () in r 4.0.3. Access information and perform operations on granges with functions from the genomicranges library. the genomation library has functions to easily load annotation files and other genomics datasets. Bondi gets news she fearedweekend update: chad maxxington on the art of looksmaxxing. Here we are going to move from iranges to granges. the key difference is that granges keeps information in "meta". the key is that the chromosome number is included in the start and stop.
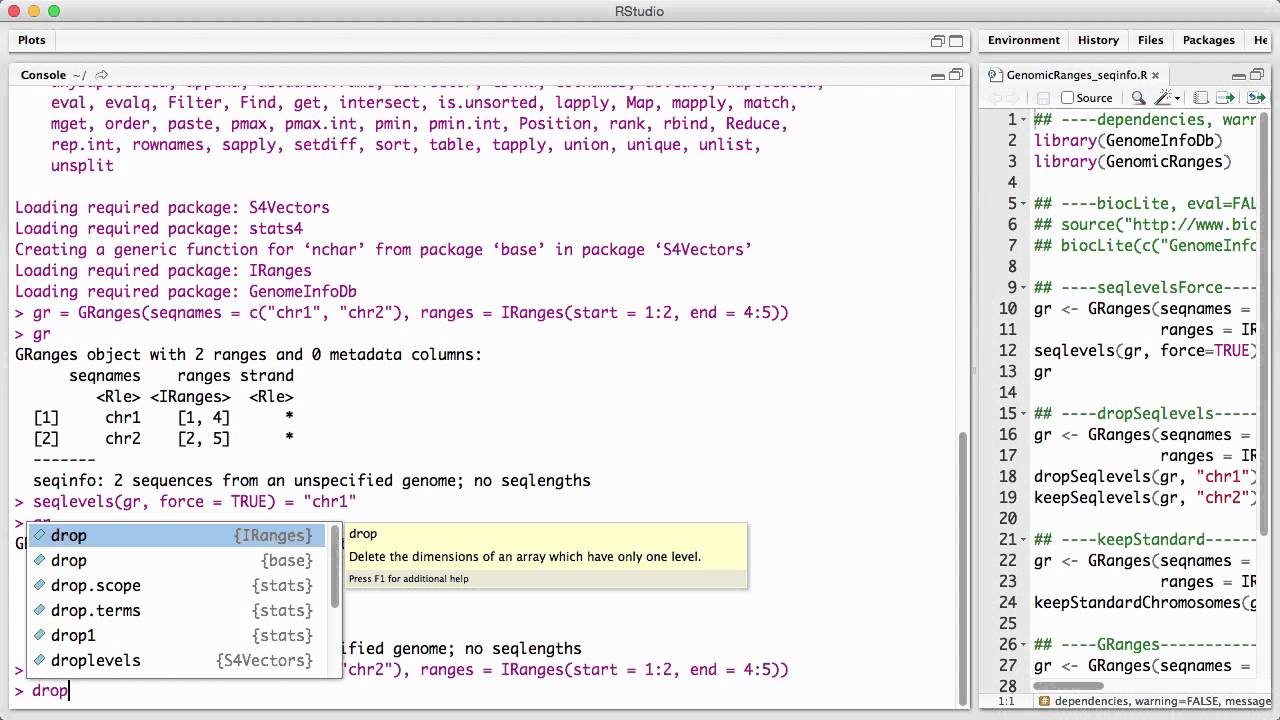
Genomicranges Seqinfo V1 Youtube Bondi gets news she fearedweekend update: chad maxxington on the art of looksmaxxing. Here we are going to move from iranges to granges. the key difference is that granges keeps information in "meta". the key is that the chromosome number is included in the start and stop.
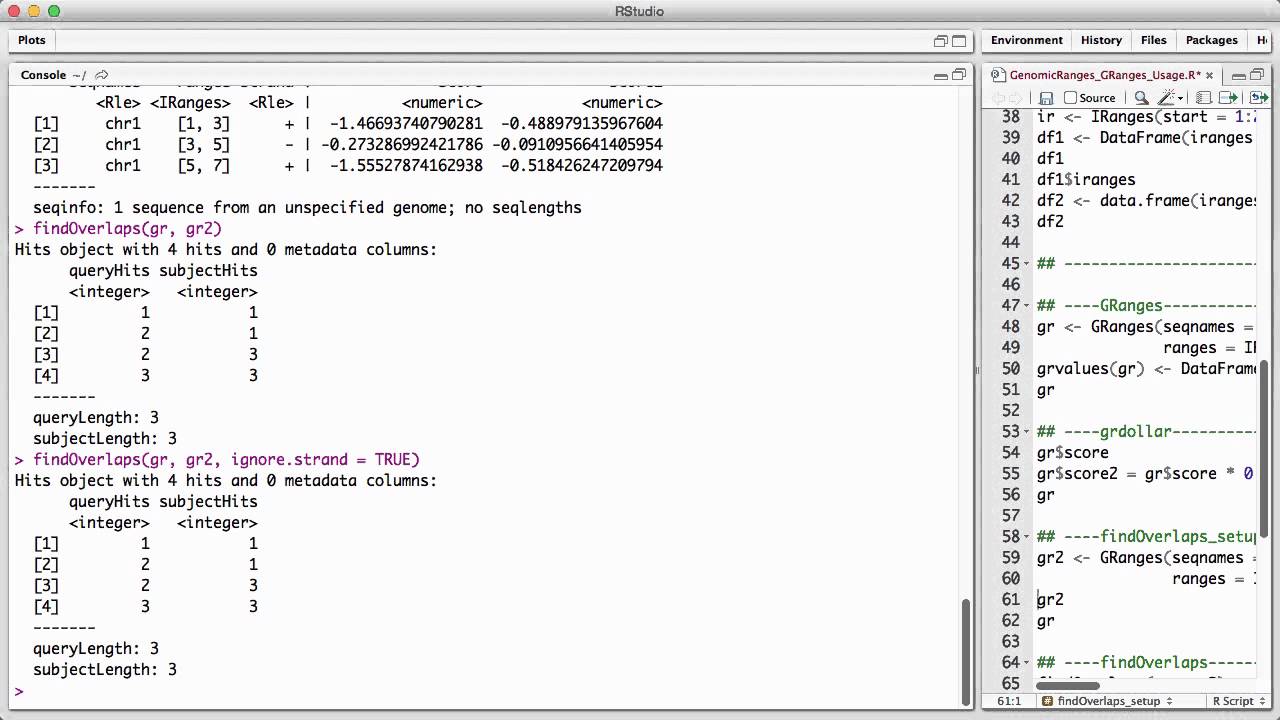
Genomicranges Granges Usage V1 Youtube
Comments are closed.