Using Annotation Packages In R For Genomic Analysis

Statistical Analysis Of Genomic Data Using R And Bioconductor Packages R packages for supporting literature reviews, data cleaning, and analyses of dna, rna, and proteins, along with their uses and strengths. Given a set of genomic sites regions (e.g. chip seq peaks, cpgs, differentially methylated cpgs or regions, snps, etc.) it is often of interest to investigate the intersecting genomic annotations.
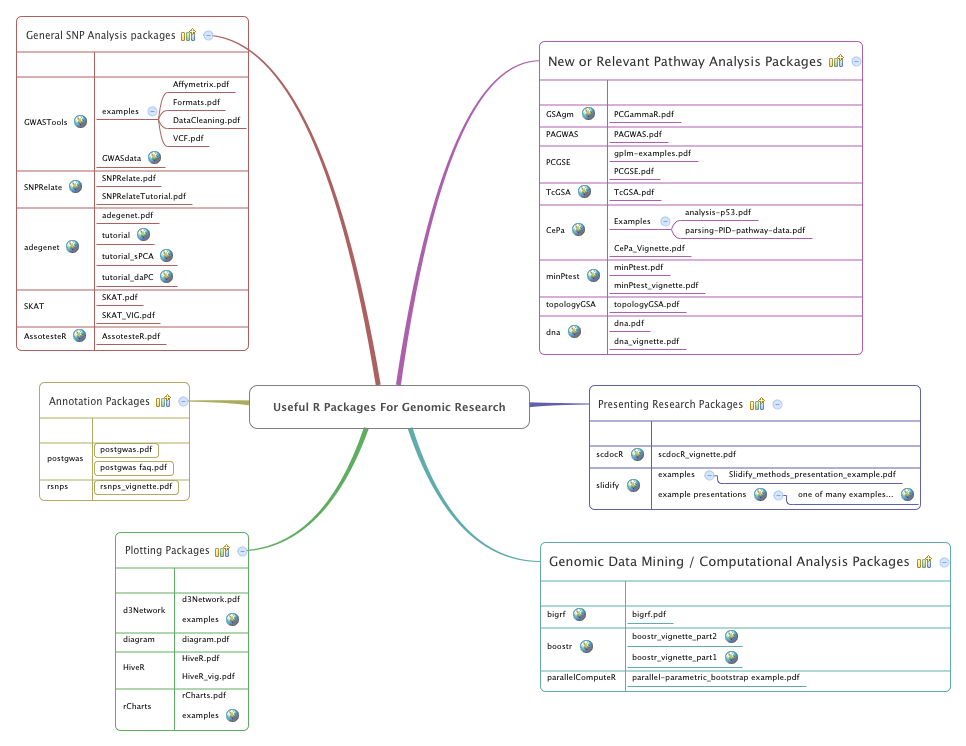
Useful R Packages For Genomic Research Masimonson Xmind A built in function easily transforms resources in the annotationhub r package (such as cosmic, encode and roadmap epigenomics) into usable annotations. finally, custom annotations can supplement built in annotations or enable annotation to any organism. Annotation data packages provide self contained databases of diverse genomic annotations (e.g., gene identifiers, biological pathways). different collections of annotation packages can be found in the bioconductor project. Given a set of genomic sites regions (e.g. chip seq peaks, cpgs, differentially methy lated cpgs or regions, snps, etc.) it is often of interest to investigate the intersecting ge nomic annotations. Packages for genomic data analysis. you can also embed plots, for example: # for working with and annotating genomes . # for analyzing high throughput sequencing data . # for handling single cell data . 3. packages for rna seq analysis. # for transcript assembly and quantification . # for visualization of rna seq data . 4.
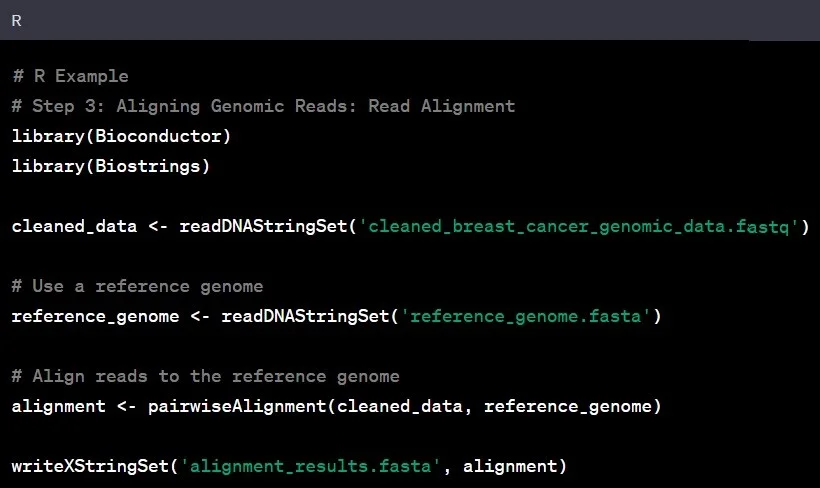
Genomic Data Analysis A Beginner S Step By Step Guide With Python And Given a set of genomic sites regions (e.g. chip seq peaks, cpgs, differentially methy lated cpgs or regions, snps, etc.) it is often of interest to investigate the intersecting ge nomic annotations. Packages for genomic data analysis. you can also embed plots, for example: # for working with and annotating genomes . # for analyzing high throughput sequencing data . # for handling single cell data . 3. packages for rna seq analysis. # for transcript assembly and quantification . # for visualization of rna seq data . 4. It allows users to visualize genome coverage with flexible input file formats, and annotate the genome coverage with various annotations to meet the needs of different ngs data. Genome centric packages are very useful for annotations involving genomic coordinates. it is straight forward, for instance, to discover the coordinates of coding sequences in regions of interest, and from these retrieve corresponding dna or protein coding sequences. In this video, i demonstrate how to utilize annotation packages in r for genomic analysis. i explain the importance of calculating total gene lengths for tasks like rpkm for rnaseq. This section details the functional annotation of genomic regions identified by cistopic. the workflow encompasses structural annotation of peaks (e.g., promoter vs. distal) using chipseeker and functional enrichment analysis of topic specific region sets using the genomic regions enrichment of annotations tool (great) via the rgreat package.
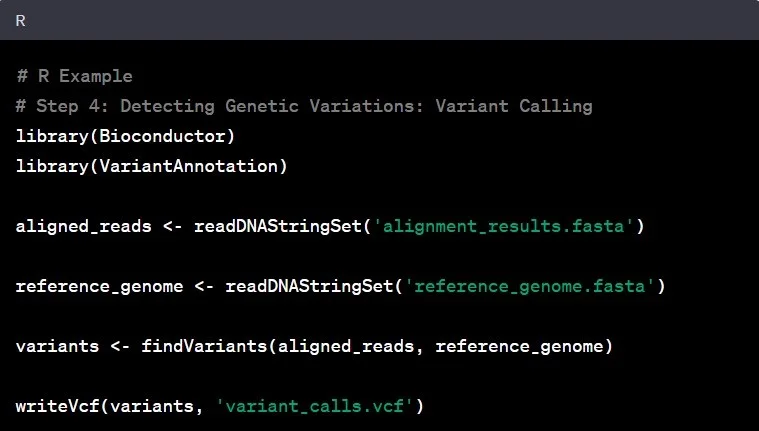
Genomic Data Analysis A Beginner S Step By Step Guide With Python And It allows users to visualize genome coverage with flexible input file formats, and annotate the genome coverage with various annotations to meet the needs of different ngs data. Genome centric packages are very useful for annotations involving genomic coordinates. it is straight forward, for instance, to discover the coordinates of coding sequences in regions of interest, and from these retrieve corresponding dna or protein coding sequences. In this video, i demonstrate how to utilize annotation packages in r for genomic analysis. i explain the importance of calculating total gene lengths for tasks like rpkm for rnaseq. This section details the functional annotation of genomic regions identified by cistopic. the workflow encompasses structural annotation of peaks (e.g., promoter vs. distal) using chipseeker and functional enrichment analysis of topic specific region sets using the genomic regions enrichment of annotations tool (great) via the rgreat package.

Shows The Results Of Annotation Analysis Using R Package Download In this video, i demonstrate how to utilize annotation packages in r for genomic analysis. i explain the importance of calculating total gene lengths for tasks like rpkm for rnaseq. This section details the functional annotation of genomic regions identified by cistopic. the workflow encompasses structural annotation of peaks (e.g., promoter vs. distal) using chipseeker and functional enrichment analysis of topic specific region sets using the genomic regions enrichment of annotations tool (great) via the rgreat package.

Shows The Results Of Annotation Analysis Using R Package Download
Comments are closed.