Single Cell Rna Seq Data Analysis Biocompiler

Single Cell Rna Seq Data Analysis Stable Diffusion Online Single cell rna sequencing is revolutionizing biology by allowing researchers to explore gene expression at the resolution of individual cells. this cutting edge technology is key to uncovering cellular diversity, understanding developmental processes, and advancing disease research. Extracting biologically interpretable insights from single cell rna sequencing (scrna seq) data remains a major challenge. while deep learning approaches have demonstrated strong predictive.
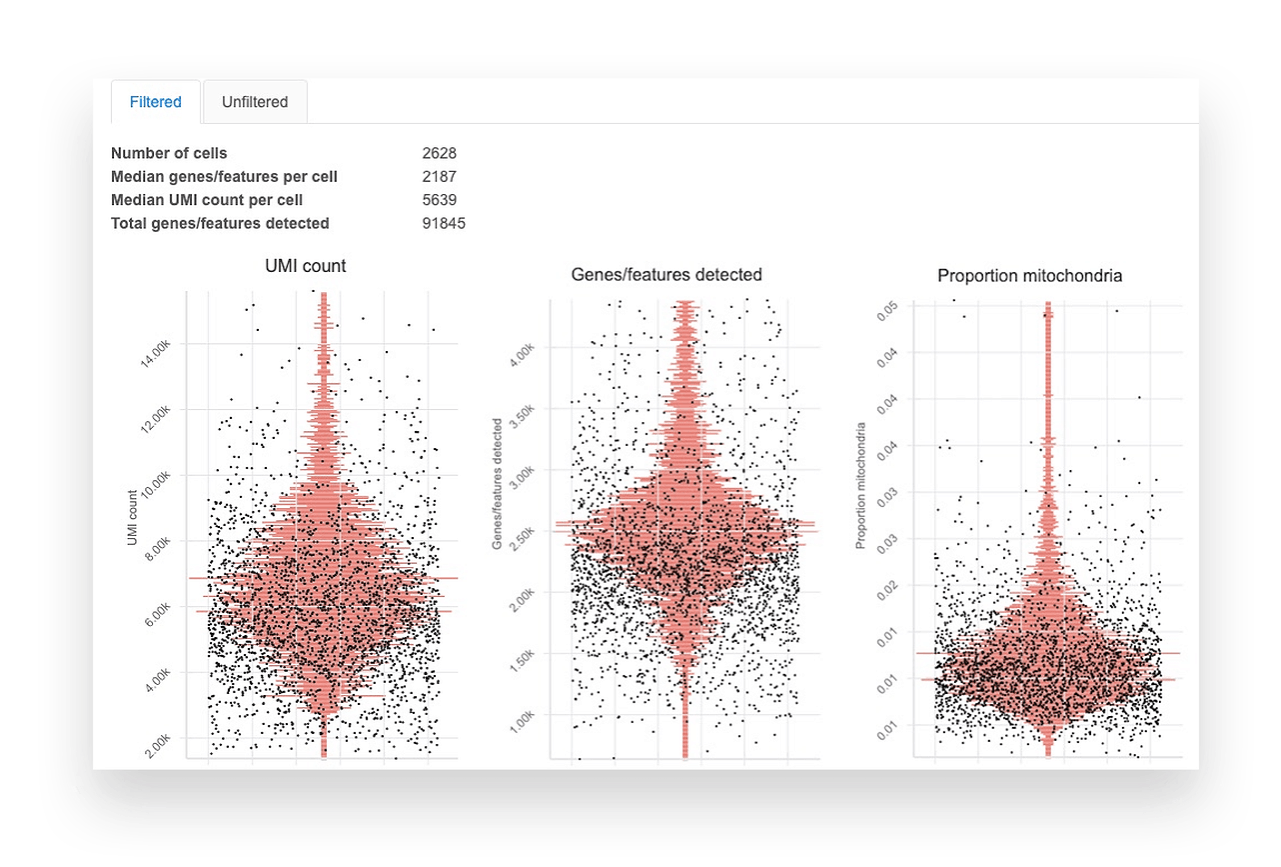
Single Cell Rna Seq Analysis Basepair In this course we will discuss some of the questions that can be addressed using scrna seq as well as the available computational and statistical methods available. This carpentries style tutorial aims to provide a solid foundation in using bioconductor tools for single cell rna seq (scrna seq) analysis by walking through various steps of typical workflows using example datasets. Background: single cell rna sequencing (scrna seq) experiments are gaining ground to study the molecular processes that drive normal development as well as the onset of different pathologies. finding an effective and efficient low dimensional representation of the data is one of the most important steps in the downstream analysis of scrna seq data, as it could provide a better identification. This dissertation develops a series of statistical and artificial intelligence–driven methodologies to improve the analysis and interpretation of single cell and gene related data. first, we introduce sciber, a simple and scalable method for correcting batch effects in scrna seq data integration.
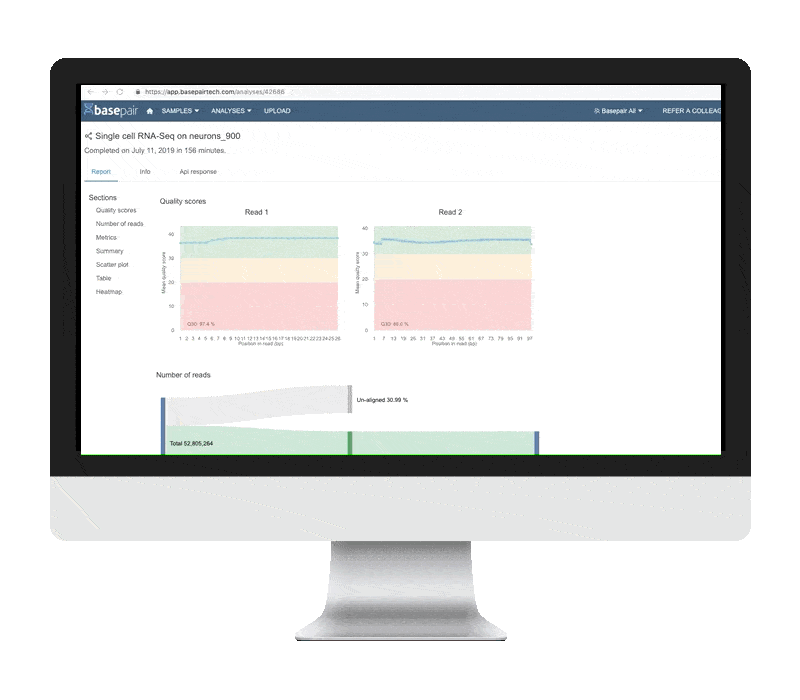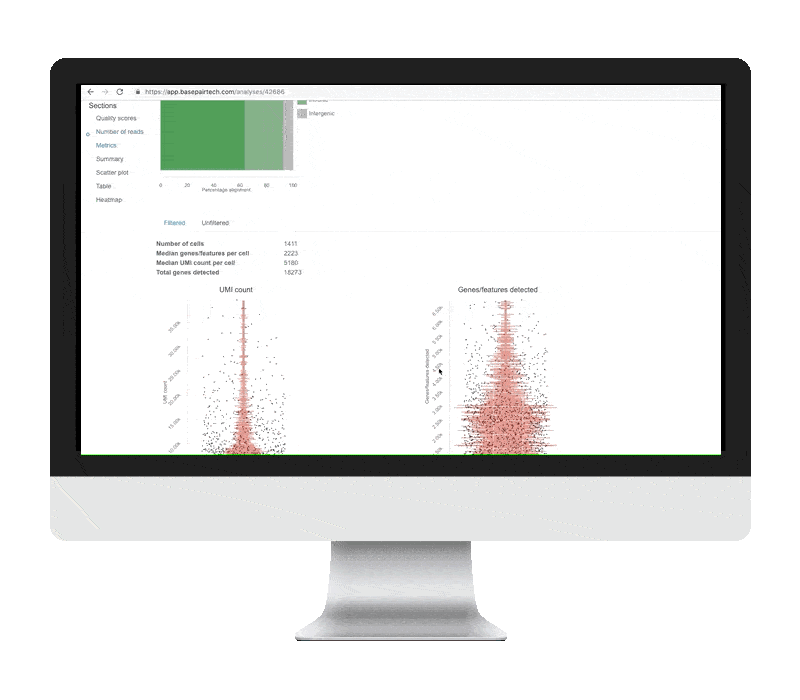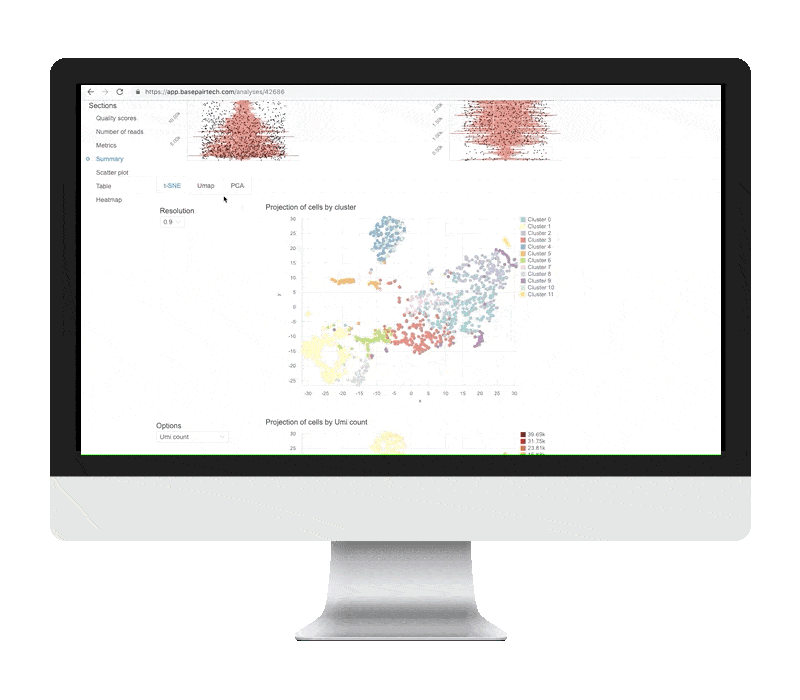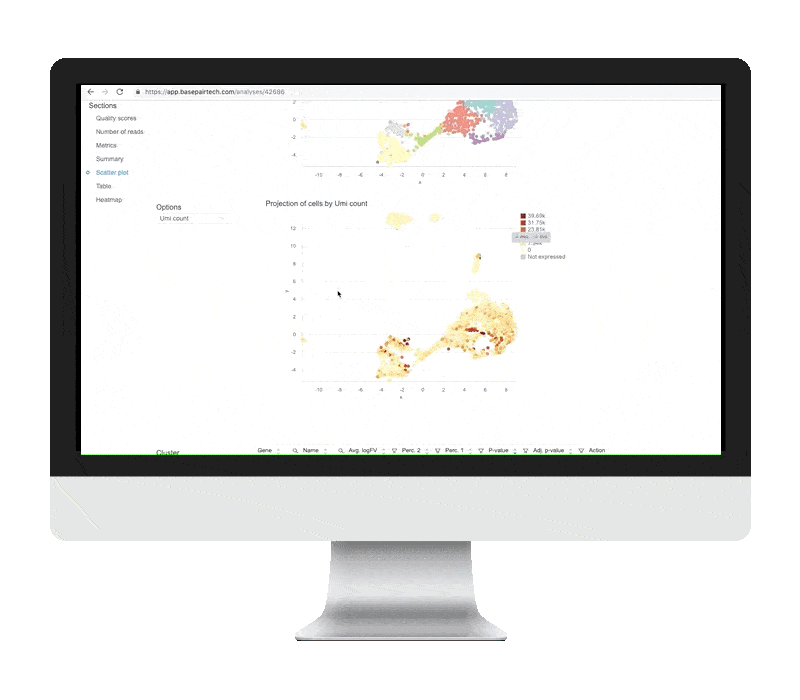
Single Cell Rna Seq Analysis Basepair Background: single cell rna sequencing (scrna seq) experiments are gaining ground to study the molecular processes that drive normal development as well as the onset of different pathologies. finding an effective and efficient low dimensional representation of the data is one of the most important steps in the downstream analysis of scrna seq data, as it could provide a better identification. This dissertation develops a series of statistical and artificial intelligence–driven methodologies to improve the analysis and interpretation of single cell and gene related data. first, we introduce sciber, a simple and scalable method for correcting batch effects in scrna seq data integration. This repository contains a demonstration workflow for single cell rna sequencing (scrna seq) analysis using a publicly available dataset of stage iii squamous cell lung carcinoma (nsclc) tumor cells. ⚠️ disclaimer: this project is intended for educational and demonstration purposes only. it is not a clinical study and does not claim any novel biological discovery. The goal of this book is to provide a solid foundation in the usage of bioconductor tools for single cell rna seq analysis by walking through various steps of typical workflows using example datasets. Single cell rna sequencing (scrna seq) enables a comprehensive analysis of the expression patterns of individual cells in tissue with the resolution of a single cell. however, “dropouts” will lead to an excess of zeros in the scrna seq data due to technical constraints, which could hinder further analysis. consequently, imputing the dropout values becomes particularly critical in assisting. Here, we review the workflow for typical scrna seq data analysis, covering raw data processing and quality control, basic data analysis applicable for almost all scrna seq data sets, and advanced data analysis that should be tailored to specific scientific questions.
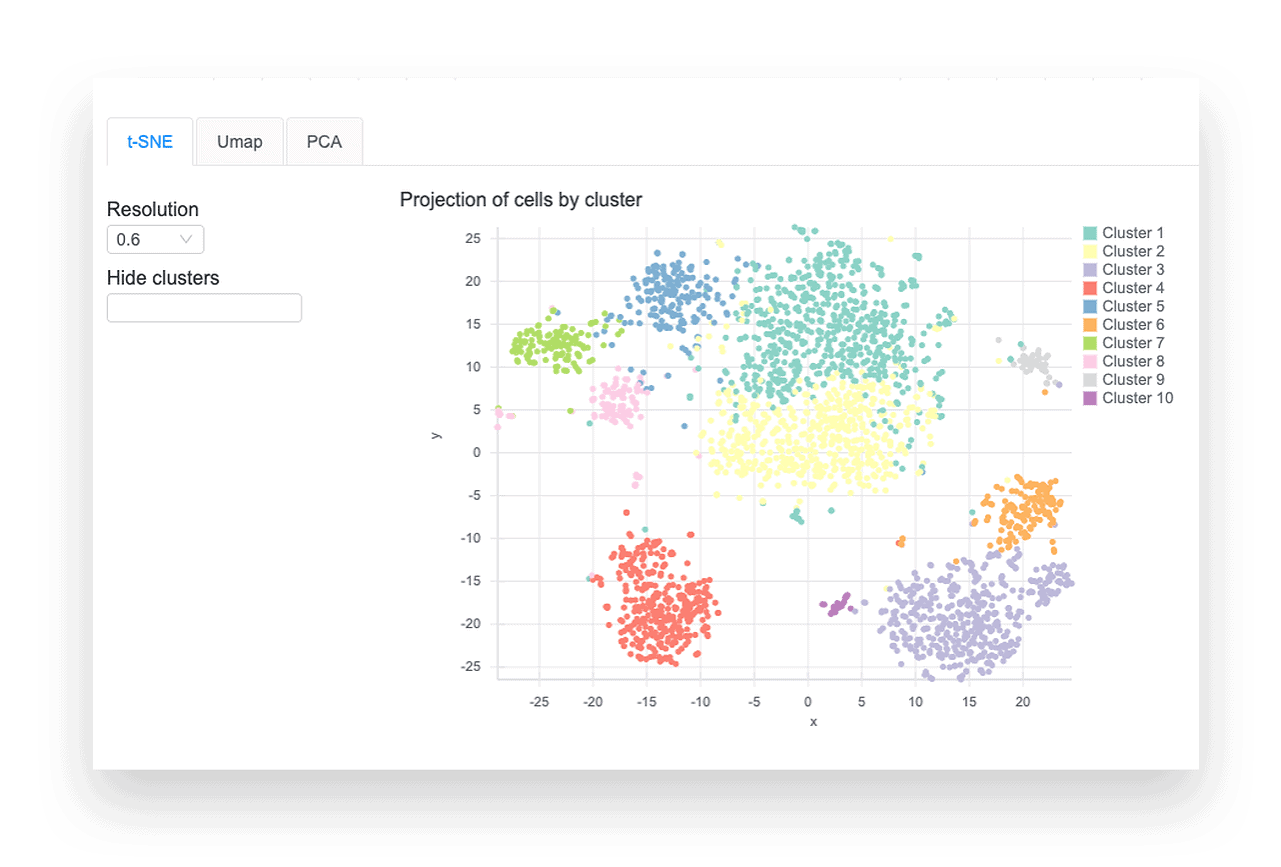
Single Cell Rna Seq Analysis Basepair This repository contains a demonstration workflow for single cell rna sequencing (scrna seq) analysis using a publicly available dataset of stage iii squamous cell lung carcinoma (nsclc) tumor cells. ⚠️ disclaimer: this project is intended for educational and demonstration purposes only. it is not a clinical study and does not claim any novel biological discovery. The goal of this book is to provide a solid foundation in the usage of bioconductor tools for single cell rna seq analysis by walking through various steps of typical workflows using example datasets. Single cell rna sequencing (scrna seq) enables a comprehensive analysis of the expression patterns of individual cells in tissue with the resolution of a single cell. however, “dropouts” will lead to an excess of zeros in the scrna seq data due to technical constraints, which could hinder further analysis. consequently, imputing the dropout values becomes particularly critical in assisting. Here, we review the workflow for typical scrna seq data analysis, covering raw data processing and quality control, basic data analysis applicable for almost all scrna seq data sets, and advanced data analysis that should be tailored to specific scientific questions.
Comments are closed.