Rplot04 Mixomics
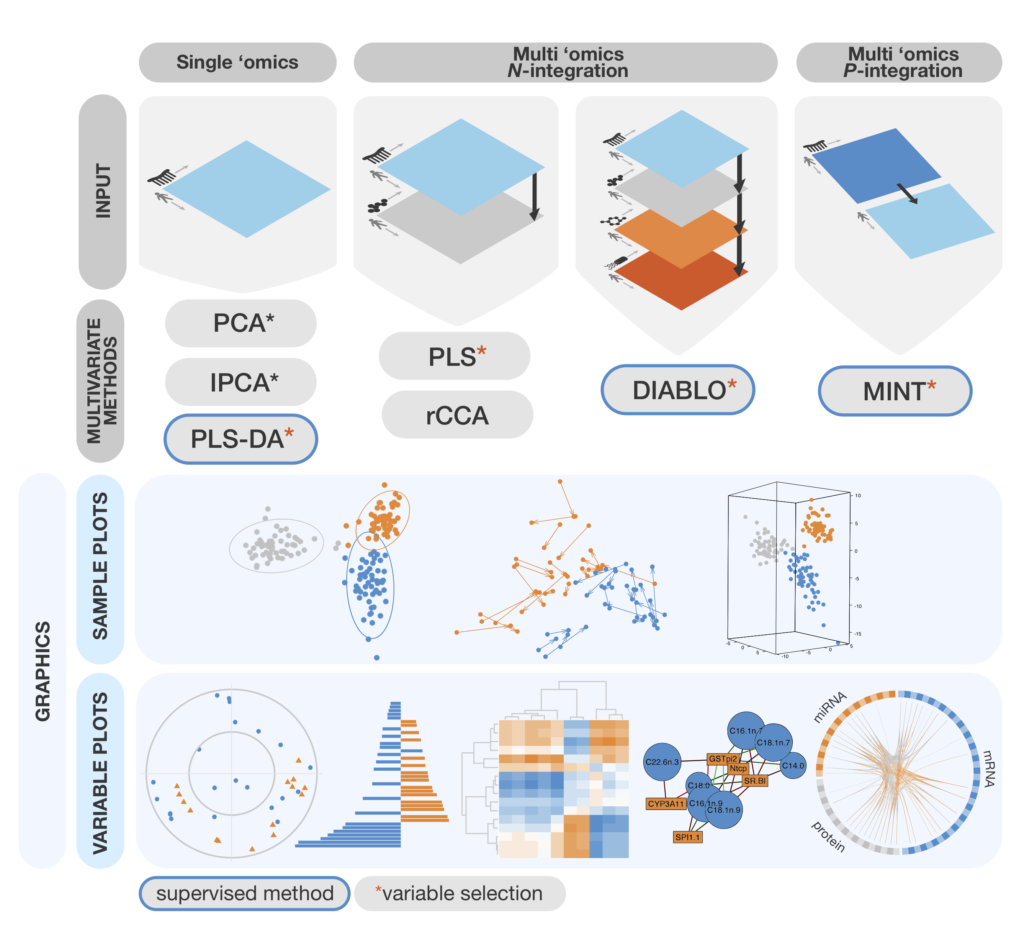
Mixomics Omics Data Integration Project The mixomics package includes tools for data integration, biomarker discovery, and data visualisation, using advanced multivariate methods to reduce data dimensionality and uncover relationships within and across datasets. Here is an overview of the most widely used methods in mixomics that will be further detailed in this vignette, with the exception of rcca. we depict them along with the type of data set they can handle.

Rplot Mixomics We introduce mixomics, an r package dedicated to the multivariate analysis of biological data sets with a specific focus on data exploration, dimension reduction and visualisation. The data are directly available in a processed and normalised format from the mixomics package and contains the following: $gene: a data frame with 63 rows and 2,308 columns. The data that can be analysed with mixomics may come from high throughput sequencing technologies, such as omics data (transcriptomics, metabolomics, proteomics, metagenomics etc) but also beyond the realm of omics (e.g. spectral imaging). We introduce mixomics in the context of supervised analysis, where the aims are to classify or discriminate sample groups, to identify the most discriminant subset of biological features, and to predict the class of new samples.

Rplot Mixomics The data that can be analysed with mixomics may come from high throughput sequencing technologies, such as omics data (transcriptomics, metabolomics, proteomics, metagenomics etc) but also beyond the realm of omics (e.g. spectral imaging). We introduce mixomics in the context of supervised analysis, where the aims are to classify or discriminate sample groups, to identify the most discriminant subset of biological features, and to predict the class of new samples. By adopting a system biology approach, the toolkit provides a wide range of methods that statistically integrate several data sets at once to probe relationships between heterogeneous ‘omics data. The data that can be analysed with mixomics may come from high throughput sequencing technologies, such as omics data (transcriptomics, metabolomics, proteomics, metagenomics etc) but also beyond the realm of omics (e.g. spectral imaging). Posted on december 4, 2015full size 832 × 574 published inrplot04. The data that can be analysed with mixomics may come from high throughput sequencing technologies, such as omics data (transcriptomics, metabolomics, proteomics, metagenomics etc) but also beyond the realm of omics (e.g. spectral imaging).

Rplot Mixomics By adopting a system biology approach, the toolkit provides a wide range of methods that statistically integrate several data sets at once to probe relationships between heterogeneous ‘omics data. The data that can be analysed with mixomics may come from high throughput sequencing technologies, such as omics data (transcriptomics, metabolomics, proteomics, metagenomics etc) but also beyond the realm of omics (e.g. spectral imaging). Posted on december 4, 2015full size 832 × 574 published inrplot04. The data that can be analysed with mixomics may come from high throughput sequencing technologies, such as omics data (transcriptomics, metabolomics, proteomics, metagenomics etc) but also beyond the realm of omics (e.g. spectral imaging).
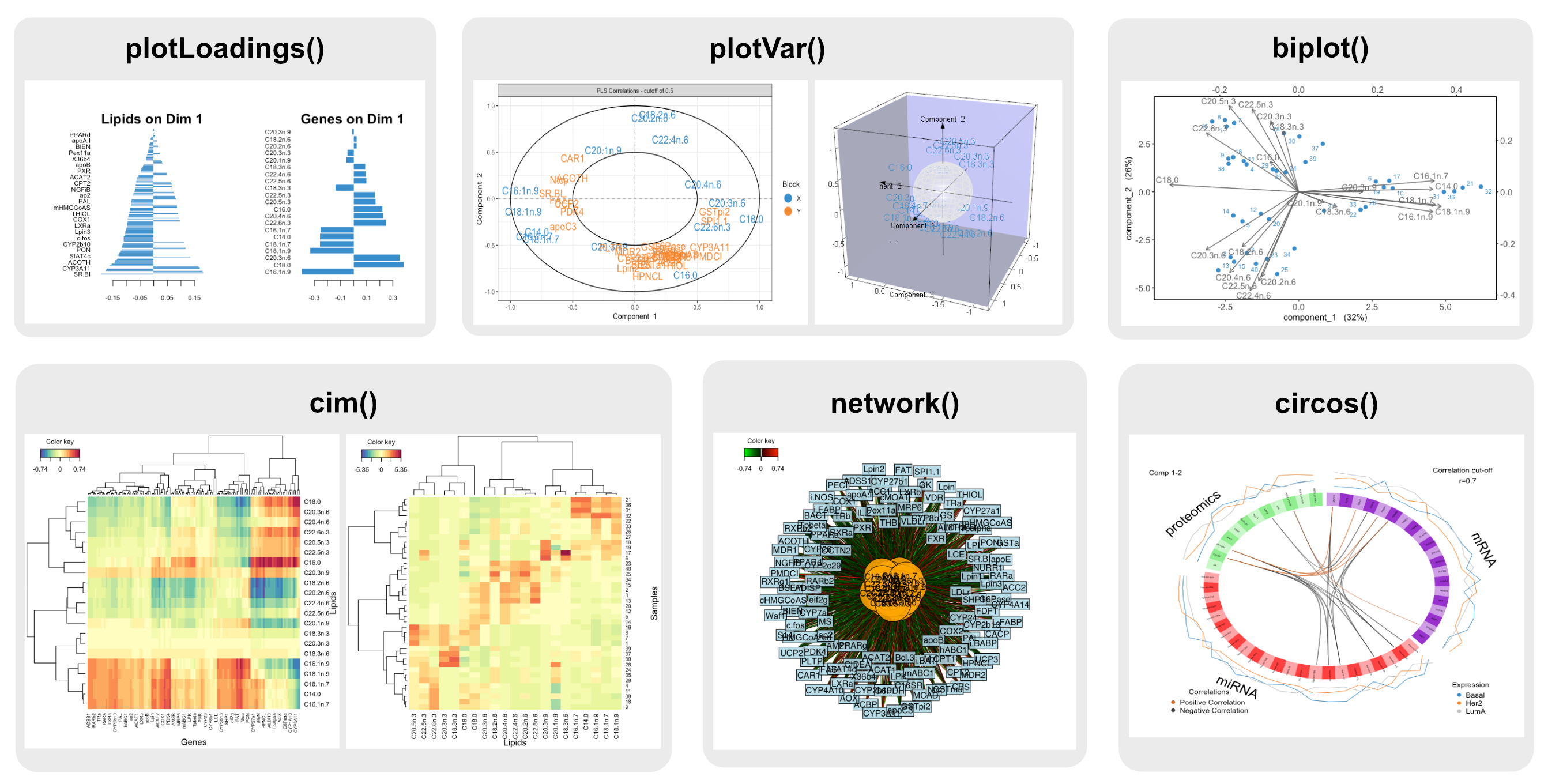
Plotting Overview Mixomics Posted on december 4, 2015full size 832 × 574 published inrplot04. The data that can be analysed with mixomics may come from high throughput sequencing technologies, such as omics data (transcriptomics, metabolomics, proteomics, metagenomics etc) but also beyond the realm of omics (e.g. spectral imaging).
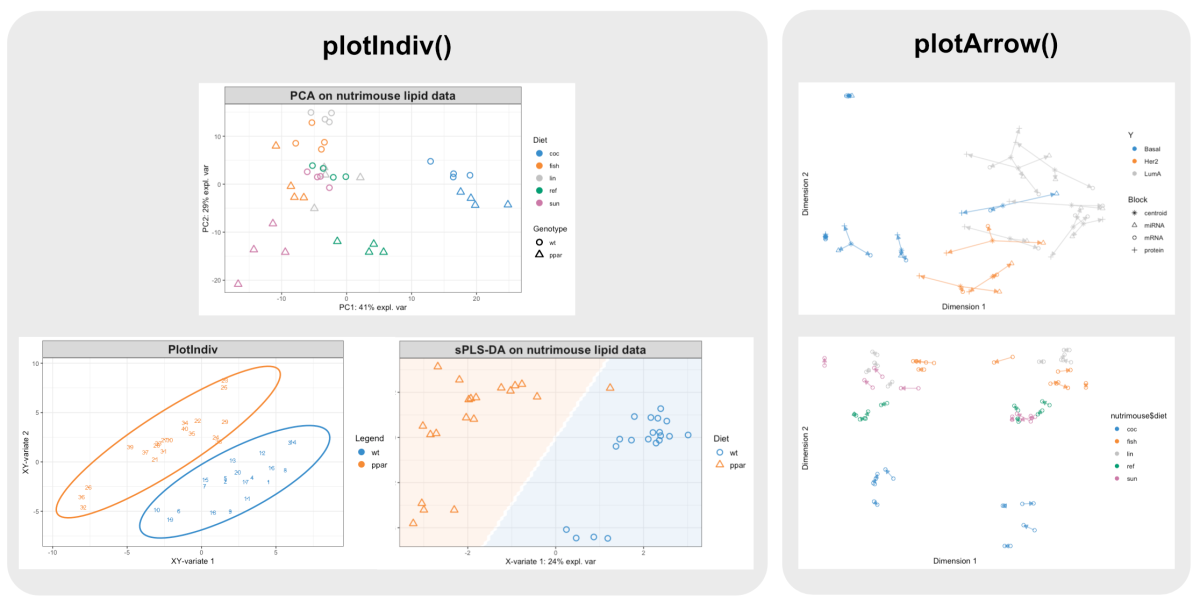
Plotting Overview Mixomics
Comments are closed.