Rplot03 Mixomics

Mixomics Pdf Principal Component Analysis Correlation And Dependence Posted on 7 december 2015full size 832 × 576 published inrplot03. You can install our latest stable github version of mixomics via our docker container. you can do this by downloading and using the docker desktop application or via the command line as described below.
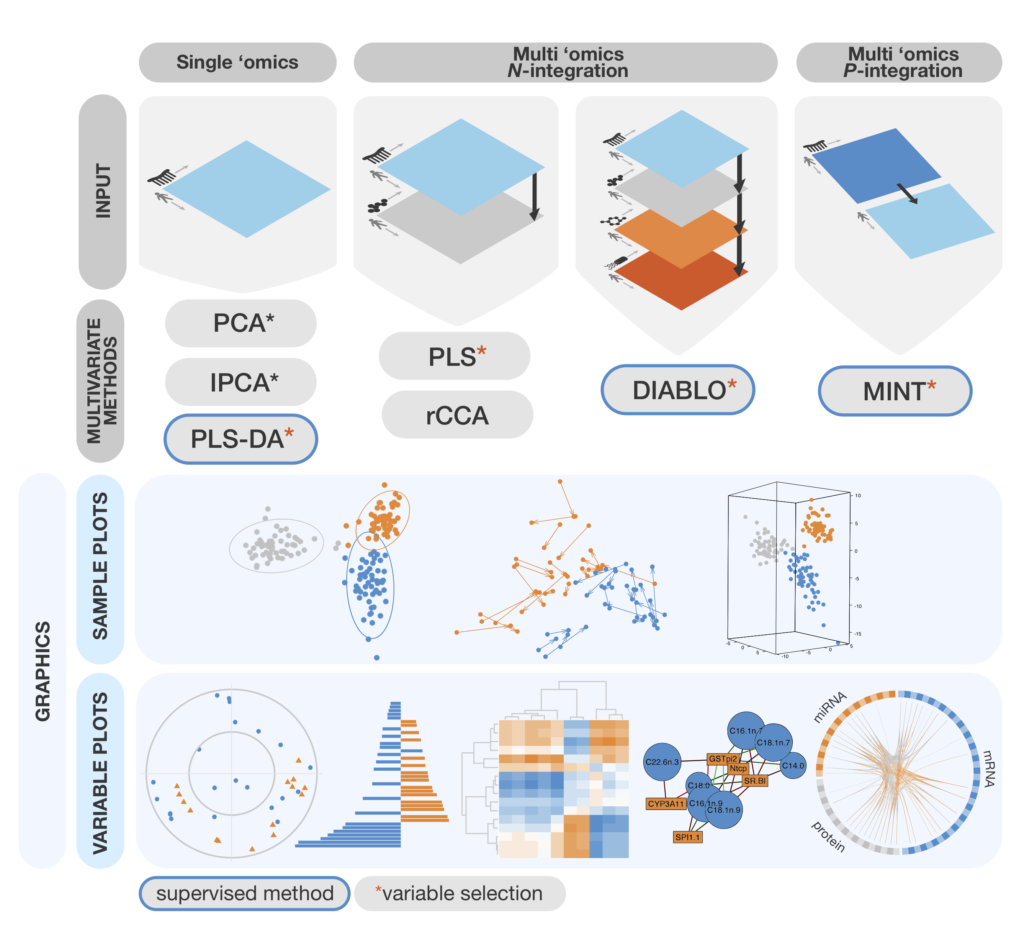
Mixomics Omics Data Integration Project The data that can be analysed with mixomics may come from high through put sequencing technologies, such as omics data (transcriptomics, metabolomics, pro teomics, metagenomics etc) but also beyond the realm of omics (e.g. spectral imag ing). Mixomics is an r package that supplies different methodologies to unravel relationships between two heterogeneous data sets of size n × p and n × q where the p and q variables are measured on the same samples or individuals n. Here is an overview of the most widely used methods in mixomics that will be further detailed in this vignette, with the exception of rcca. we depict them along with the type of data set they can handle. We introduce mixomics, an r package dedicated to the multivariate analysis of biological data sets with a specific focus on data exploration, dimension reduction and visualisation.

Rplot Mixomics Here is an overview of the most widely used methods in mixomics that will be further detailed in this vignette, with the exception of rcca. we depict them along with the type of data set they can handle. We introduce mixomics, an r package dedicated to the multivariate analysis of biological data sets with a specific focus on data exploration, dimension reduction and visualisation. We introduce mixomics in the context of supervised analysis, where the aims are to classify or discriminate sample groups, to identify the most discriminant subset of biological features, and to predict the class of new samples. The data that can be analysed with mixomics may come from high throughput sequencing technologies, such as omics data (transcriptomics, metabolomics, proteomics, metagenomics etc) but also beyond the realm of omics (e.g. spectral imaging). The data that can be analysed with mixomics may come from high throughput sequencing technologies, such as omics data (transcriptomics, metabolomics, proteomics, metagenomics etc) but also beyond the realm of omics (e.g. spectral imaging). The data that can be analysed with mixomics may come from high throughput sequencing technologies, such as omics data (transcriptomics, metabolomics, proteomics, metagenomics etc) but also beyond the realm of omics (e.g. spectral imaging).

Rplot Mixomics We introduce mixomics in the context of supervised analysis, where the aims are to classify or discriminate sample groups, to identify the most discriminant subset of biological features, and to predict the class of new samples. The data that can be analysed with mixomics may come from high throughput sequencing technologies, such as omics data (transcriptomics, metabolomics, proteomics, metagenomics etc) but also beyond the realm of omics (e.g. spectral imaging). The data that can be analysed with mixomics may come from high throughput sequencing technologies, such as omics data (transcriptomics, metabolomics, proteomics, metagenomics etc) but also beyond the realm of omics (e.g. spectral imaging). The data that can be analysed with mixomics may come from high throughput sequencing technologies, such as omics data (transcriptomics, metabolomics, proteomics, metagenomics etc) but also beyond the realm of omics (e.g. spectral imaging).
Comments are closed.