Rnaseq Data Preprocessing Using Shell Commands In Linux
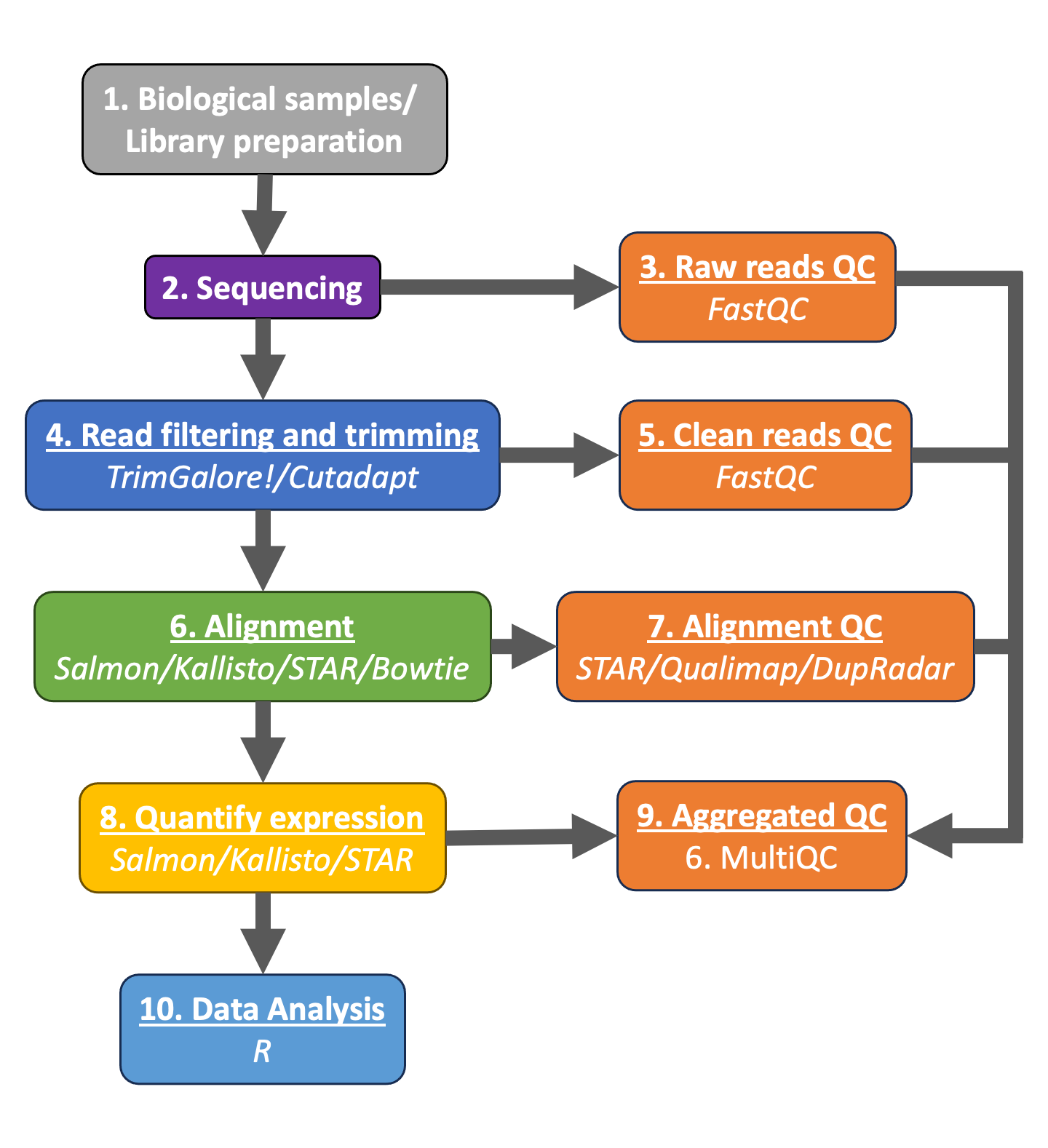
Preprocessing Steps Bulk Rnaseq Data Analysis The raw data of any ngs experiment is provided in a file format named fastq, handling such files is essential for any bioinformatician. to achieve this task, shell commands in a linux system are commonly used. This involves using shell commands in a linux environment.in this module, you'll become acquainted with the linux system and shell commands, learn about various data file formats, and discover how to select and retrieve raw files from the gene expression omnibus (geo).
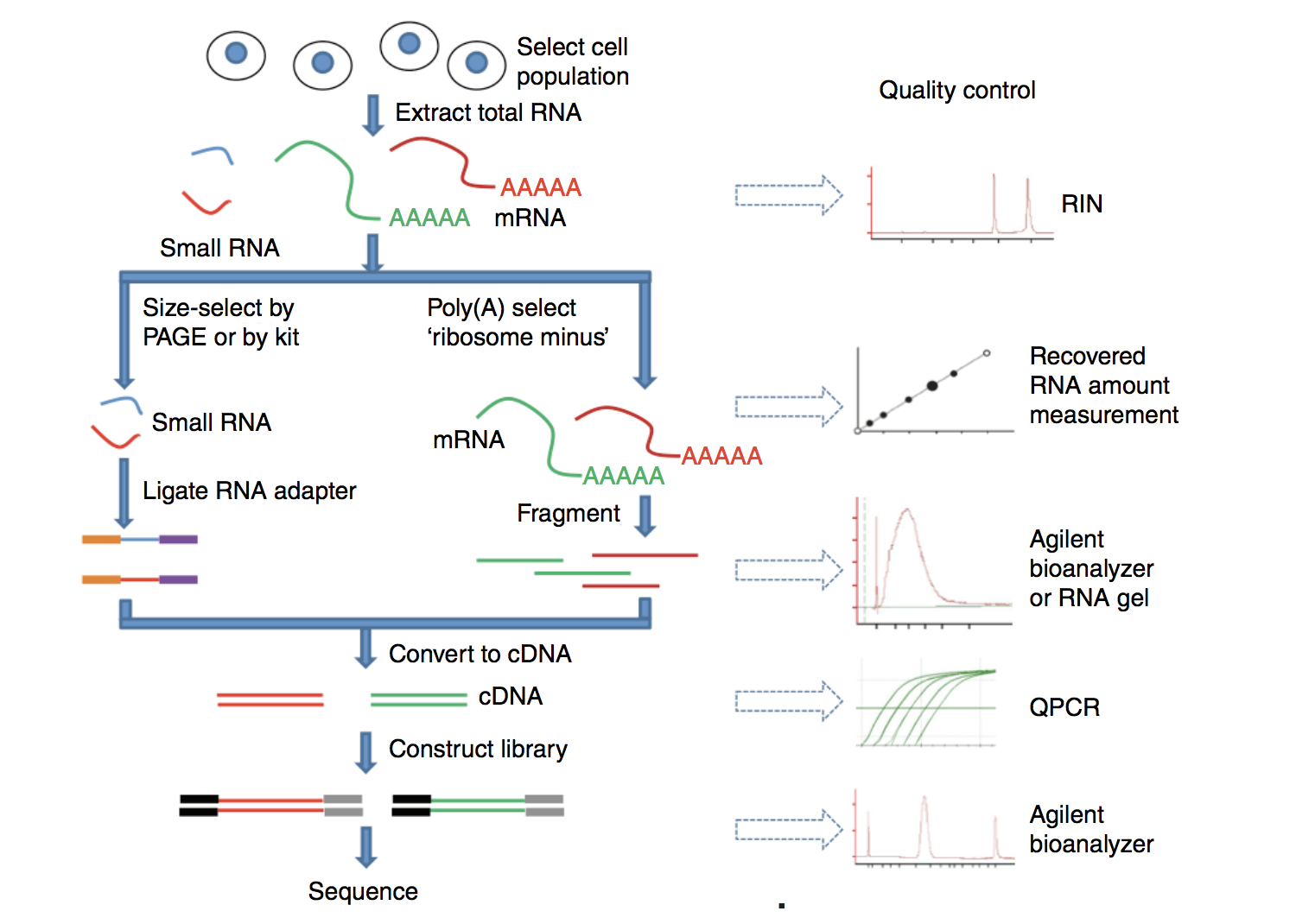
Preprocessing Steps Bulk Rnaseq Data Analysis In this course, you will learn how to perform rnaseq data analysis via linux command line. this course provides a comprehensive introduction to rnaseq data analysis, covering the key concepts and tools needed to perform differential expression analysis and functional annotation of rnaseq data. We are using the linux command line to run most of the tools we use today. if you are new to linux please complete the intro to command line workshop. for a quick refresh of linux commands take a look here. we will use the wcb bioinformatics servers for this practical. This repository provides a streamlined and automated shell script for preprocessing raw rna seq reads. it is built using a robust combination of fastqc for quality control and fastp for high performance adapter and quality trimming. Learn data science & ai with rnaseq data analysis using shell scripting and r. in this course, you will learn how to perform rnaseq data analysis via linux command line.

Rnaseq Data Analysis Using Shell Scripting And R Softarchive This repository provides a streamlined and automated shell script for preprocessing raw rna seq reads. it is built using a robust combination of fastqc for quality control and fastp for high performance adapter and quality trimming. Learn data science & ai with rnaseq data analysis using shell scripting and r. in this course, you will learn how to perform rnaseq data analysis via linux command line. Commands in bash are executed through scripts, which are delineated with a “.sh” at the end. think of them as packages containing instructions that tell your computer (or hipergator) what to do. You must also have familiarity with using the command line to interact with linux systems. the tutorial will use standard linux utilities to navigate the file system, and our rnaseq tools are written to work from the command line. The first part of this computational pipeline is performed in terminal, also known as shell, which runs code from the command line, ideally using a high capacity computer for faster data processing. In this course, you will learn how to perform rnaseq data analysis via linux command line. this course provides a comprehensive introduction to rnaseq data analysis, covering the key concepts and tools needed to perform differential expression analysis and functional annotation of rnaseq data.
Rnaseq Data Preprocessing Using Shell Commands In Linux Commands in bash are executed through scripts, which are delineated with a “.sh” at the end. think of them as packages containing instructions that tell your computer (or hipergator) what to do. You must also have familiarity with using the command line to interact with linux systems. the tutorial will use standard linux utilities to navigate the file system, and our rnaseq tools are written to work from the command line. The first part of this computational pipeline is performed in terminal, also known as shell, which runs code from the command line, ideally using a high capacity computer for faster data processing. In this course, you will learn how to perform rnaseq data analysis via linux command line. this course provides a comprehensive introduction to rnaseq data analysis, covering the key concepts and tools needed to perform differential expression analysis and functional annotation of rnaseq data.
Github Irishevia Rnaseq Box Dockerized Rna Seq Analysis Pipeline The first part of this computational pipeline is performed in terminal, also known as shell, which runs code from the command line, ideally using a high capacity computer for faster data processing. In this course, you will learn how to perform rnaseq data analysis via linux command line. this course provides a comprehensive introduction to rnaseq data analysis, covering the key concepts and tools needed to perform differential expression analysis and functional annotation of rnaseq data.

Rnaseq Preprocessing And Analysis Tools Currently In Common Use
Comments are closed.