Reliable Nuclei Quality Assessment For Single Cell Genomics

Advanced Nuclei Quality Assessment In Single Cell Genomics This study aims to identify the most effective dye combinations for nuclei quality assessment using different cell types, different systems, and nuclei fixation. Here, we provide the groundwork for improving the quality of single cell analysis by delineating guidelines for selecting high quality cells and considerations throughout the analysis. this review will streamline researchers' access to single cell analysis and serve as a valuable guide for analysis.

Advanced Nuclei Quality Assessment In Single Cell Genomics In this guide, we provide an overview of best practices for analyzing single cell gene expression data generated using the chromium platform from 10x genomics, along with an example experimental setup for multiple datasets. Quality control (qc) of cells, a critical first step in single cell rna sequencing data analysis, has largely relied on arbitrarily fixed data agnostic thresholds applied to qc metrics such as gene complexity and fraction of reads mapping to mitochondrial genes. We showcase how each step in validrops improves the data quality and we extensively benchmark validrops and existing methods using 47 real samples from five different studies to show that validrops has the best performance for both single cell and single nucleus rna seq data. Single cell rna seq datasets have two essential properties that one should have in mind when performing an analysis. first, scrna seq data contains drop outs, meaning there is an excessive number of zeros in the data due to the limitation of mrna.
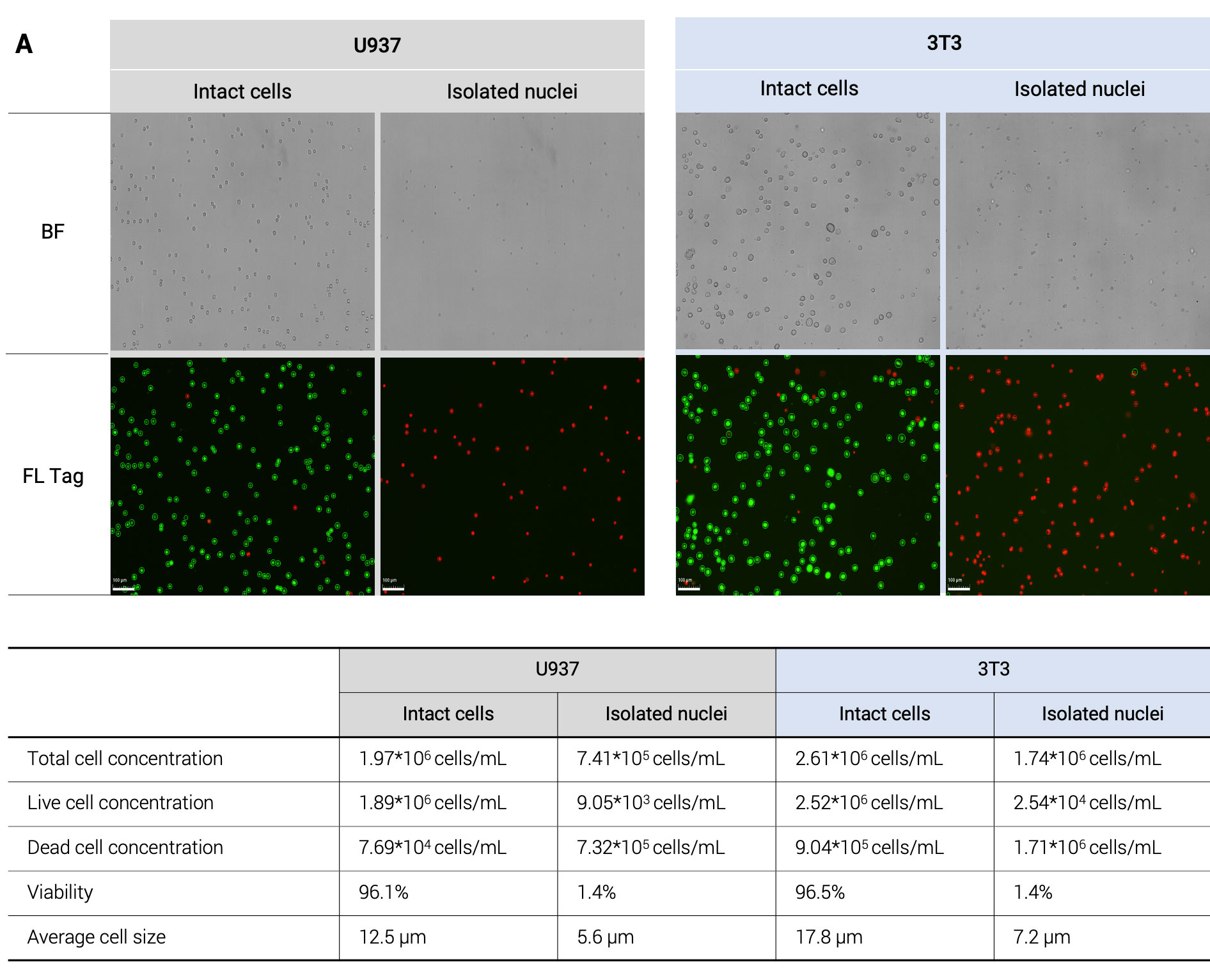
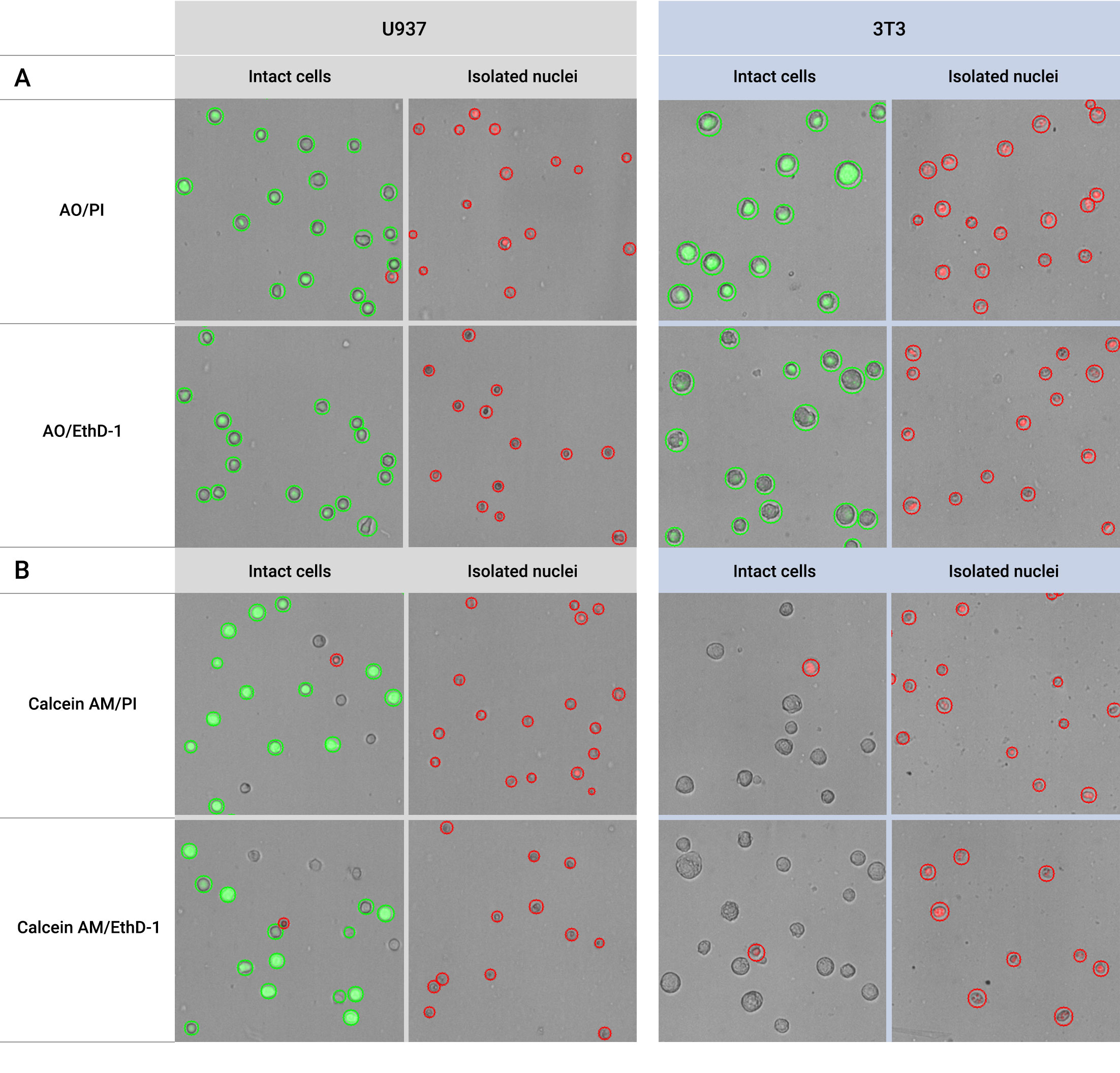
Reliable Nuclei Quality Assessment For Single Cell Genomics We showcase how each step in validrops improves the data quality and we extensively benchmark validrops and existing methods using 47 real samples from five different studies to show that validrops has the best performance for both single cell and single nucleus rna seq data. Single cell rna seq datasets have two essential properties that one should have in mind when performing an analysis. first, scrna seq data contains drop outs, meaning there is an excessive number of zeros in the data due to the limitation of mrna. We use several common qc metrics to identify low quality cells based on their expression profiles. these metrics are described below in terms of reads for smart seq2 data, but the same definitions apply to umi data generated by other technologies like mars seq and droplet based protocols. In this study, we use intronic fraction and malat1 expression to assess cell quality in several publicly available datasets from reference atlases. The accurate evaluation of nuclei quality is pivotal for single cell genomics research. combining fluorescent dyes with imaging tools such as automated cell counters, provides an effective method to evaluate nuclei quality. Single cell rna sequencing (scrna seq) can be used to gain insights into cellular heterogeneity within complex tissues. however, various technical artifacts can be present in scrna seq data and.

10x Genomics Single Cell Nuclei Spatial Technologies Unit We use several common qc metrics to identify low quality cells based on their expression profiles. these metrics are described below in terms of reads for smart seq2 data, but the same definitions apply to umi data generated by other technologies like mars seq and droplet based protocols. In this study, we use intronic fraction and malat1 expression to assess cell quality in several publicly available datasets from reference atlases. The accurate evaluation of nuclei quality is pivotal for single cell genomics research. combining fluorescent dyes with imaging tools such as automated cell counters, provides an effective method to evaluate nuclei quality. Single cell rna sequencing (scrna seq) can be used to gain insights into cellular heterogeneity within complex tissues. however, various technical artifacts can be present in scrna seq data and.
Comments are closed.