Pygengraph User Manual
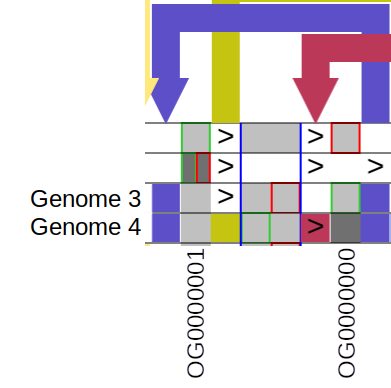
Pygengraph User Manual This manual describe how to use all the controls and how to read the matrix view of the graph visualisation. it will cover two types of graphs: nucleotide and gene graphs. Python package for generation construction and processing of genome graphs; includes exporting for visualisation in pantograph visualisation tool. pygengraph user manual.md at master · pigrenok pygengraph.

Pygengraph User Manual User manual for sonargraph software v9.9.0, covering installation, licensing, system model, module creation, and interaction. Find instructions for use, manuals, dicom and ihe statements and more on our document resource center. The rest of the file describes some of the uses of the pygengraph package. there are more ways to use it, but more detailed documentation is needed to describe all use cases. The rest of the file describes some of the uses of the pygengraph package. there are more ways to use it, but more detailed documentation is needed to describe all use cases.

Pygengraph User Manual The rest of the file describes some of the uses of the pygengraph package. there are more ways to use it, but more detailed documentation is needed to describe all use cases. The rest of the file describes some of the uses of the pygengraph package. there are more ways to use it, but more detailed documentation is needed to describe all use cases. Chromosome: str or none, if none, create one graph for all chromosomes (not fully implemented, see manual), otherwise, create only one graph for given chromosome. Credit: implemented as an answer to stackoverflow questions 3318625 how to implement an efficient bidirectional hash table by basj ( stackoverflow users 1422096 basj). {"payload":{"allshortcutsenabled":false,"filetree":{"":{"items":[{"name":".github","path":".github","contenttype":"directory"},{"name":"docs","path":"docs","contenttype":"directory"},{"name":"examples","path":"examples","contenttype":"directory"},{"name":"images","path":"images","contenttype":"directory"},{"name":"pygengraph","path":"pygengraph","contenttype":"directory"},{"name":".devcontainer.json","path":".devcontainer.json","contenttype":"file"},{"name":".gitignore","path":".gitignore","contenttype":"file"},{"name":".gitlab ci.yml","path":".gitlab ci.yml","contenttype":"file"},{"name":"00 init.ipynb","path":"00 init.ipynb","contenttype":"file"},{"name":"01 graph.ipynb","path":"01 graph.ipynb","contenttype":"file"},{"name":"02 tree.ipynb","path":"02 tree.ipynb","contenttype":"file"},{"name":"03 synteny.ipynb","path":"03 synteny.ipynb","contenttype":"file"},{"name":"04 utils.ipynb","path":"04 utils.ipynb","contenttype":"file"},{"name":"05 export.ipynb","path":"05 export.ipynb","contenttype":"file"},{"name":"06. Provides functionality to construct special synteny graph from pangenome annotation (and actual sequences).
Pygengraph User Manual Chromosome: str or none, if none, create one graph for all chromosomes (not fully implemented, see manual), otherwise, create only one graph for given chromosome. Credit: implemented as an answer to stackoverflow questions 3318625 how to implement an efficient bidirectional hash table by basj ( stackoverflow users 1422096 basj). {"payload":{"allshortcutsenabled":false,"filetree":{"":{"items":[{"name":".github","path":".github","contenttype":"directory"},{"name":"docs","path":"docs","contenttype":"directory"},{"name":"examples","path":"examples","contenttype":"directory"},{"name":"images","path":"images","contenttype":"directory"},{"name":"pygengraph","path":"pygengraph","contenttype":"directory"},{"name":".devcontainer.json","path":".devcontainer.json","contenttype":"file"},{"name":".gitignore","path":".gitignore","contenttype":"file"},{"name":".gitlab ci.yml","path":".gitlab ci.yml","contenttype":"file"},{"name":"00 init.ipynb","path":"00 init.ipynb","contenttype":"file"},{"name":"01 graph.ipynb","path":"01 graph.ipynb","contenttype":"file"},{"name":"02 tree.ipynb","path":"02 tree.ipynb","contenttype":"file"},{"name":"03 synteny.ipynb","path":"03 synteny.ipynb","contenttype":"file"},{"name":"04 utils.ipynb","path":"04 utils.ipynb","contenttype":"file"},{"name":"05 export.ipynb","path":"05 export.ipynb","contenttype":"file"},{"name":"06. Provides functionality to construct special synteny graph from pangenome annotation (and actual sequences).
Github Blantagoose Manualphylogeny Script For Creating Phylogeny By {"payload":{"allshortcutsenabled":false,"filetree":{"":{"items":[{"name":".github","path":".github","contenttype":"directory"},{"name":"docs","path":"docs","contenttype":"directory"},{"name":"examples","path":"examples","contenttype":"directory"},{"name":"images","path":"images","contenttype":"directory"},{"name":"pygengraph","path":"pygengraph","contenttype":"directory"},{"name":".devcontainer.json","path":".devcontainer.json","contenttype":"file"},{"name":".gitignore","path":".gitignore","contenttype":"file"},{"name":".gitlab ci.yml","path":".gitlab ci.yml","contenttype":"file"},{"name":"00 init.ipynb","path":"00 init.ipynb","contenttype":"file"},{"name":"01 graph.ipynb","path":"01 graph.ipynb","contenttype":"file"},{"name":"02 tree.ipynb","path":"02 tree.ipynb","contenttype":"file"},{"name":"03 synteny.ipynb","path":"03 synteny.ipynb","contenttype":"file"},{"name":"04 utils.ipynb","path":"04 utils.ipynb","contenttype":"file"},{"name":"05 export.ipynb","path":"05 export.ipynb","contenttype":"file"},{"name":"06. Provides functionality to construct special synteny graph from pangenome annotation (and actual sequences).
Pangraphviewer Doc Manual Pdf At Main Tf Chan Lab Pangraphviewer Github
Comments are closed.