Ngs Pipeline Github Topics Github
Ngs Pipeline Github Topics Github Add a description, image, and links to the ngs pipeline topic page so that developers can more easily learn about it. to associate your repository with the ngs pipeline topic, visit your repo's landing page and select "manage topics." github is where people build software. Ngspipedb is an automated pipeline for parallel processing of huge next generation sequencing (ngs) data and database generation using snakemake workflow which allows for ease of use, optimal speed, and a highly modular code that can be further added onto and customized by experienced users.
Activity Ipseity Bio Ngs Pipeline Github Genome assembly • apply analysis pipelines to generate high quality genome assemblies many different strategies, implementations & can be used! • a challenge for ngi scilifelab is to give best practice guidelines!. There are a number of pipelines available both commerical and academic, with some that are specific to a particular ngs experiment (i.e variant calling, rna seq, viral ngs analysis). the pipeline we will be presenting here is bcbio nextgen. In this talk, we’ll cover concepts & tips for making the most of github to manage bioinformatics projects, and we’ll demonstrate how we use these in practice with ccbr pipelines. In this tutorial you will bring a set of fastq files representing reads from an illumina sequencing platform through a basic ngs data processing pipeline. you will have to qc and align the reads, identify or remove duplicate reads, compute coverage and generate an initial set of variant calls.
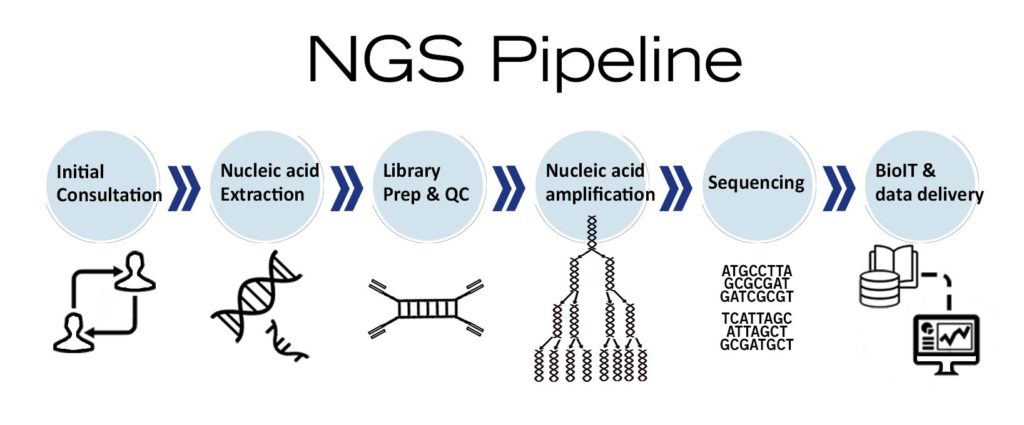
Ngs A Powerful Tool For Disease Diagnostics In this talk, we’ll cover concepts & tips for making the most of github to manage bioinformatics projects, and we’ll demonstrate how we use these in practice with ccbr pipelines. In this tutorial you will bring a set of fastq files representing reads from an illumina sequencing platform through a basic ngs data processing pipeline. you will have to qc and align the reads, identify or remove duplicate reads, compute coverage and generate an initial set of variant calls. Here, we introduce ngs pipe, an automated and user friendly framework for the design of pipelines for the analysis of large scale sequencing data, such as cancer genomics data. A bioinformatics best practice analysis pipeline for calling structural variants (svs), copy number variants (cnvs) and repeat region expansions (rres) from short dna reads. In summary, we believe that neat will help biologists as well as established bioinformaticians create, manage and analyze complex ngs pipelines, as well as assess ngs data within 24 h of the sequencing run completion through a simple gui. Customizable workflows based on snakemake and python for the analysis of ngs data.
Github Abcsfrederick Ngs Preprocessing Pipeline Ngs Pipelines For Here, we introduce ngs pipe, an automated and user friendly framework for the design of pipelines for the analysis of large scale sequencing data, such as cancer genomics data. A bioinformatics best practice analysis pipeline for calling structural variants (svs), copy number variants (cnvs) and repeat region expansions (rres) from short dna reads. In summary, we believe that neat will help biologists as well as established bioinformaticians create, manage and analyze complex ngs pipelines, as well as assess ngs data within 24 h of the sequencing run completion through a simple gui. Customizable workflows based on snakemake and python for the analysis of ngs data.
Comments are closed.