Molecular Dynamics Simulations
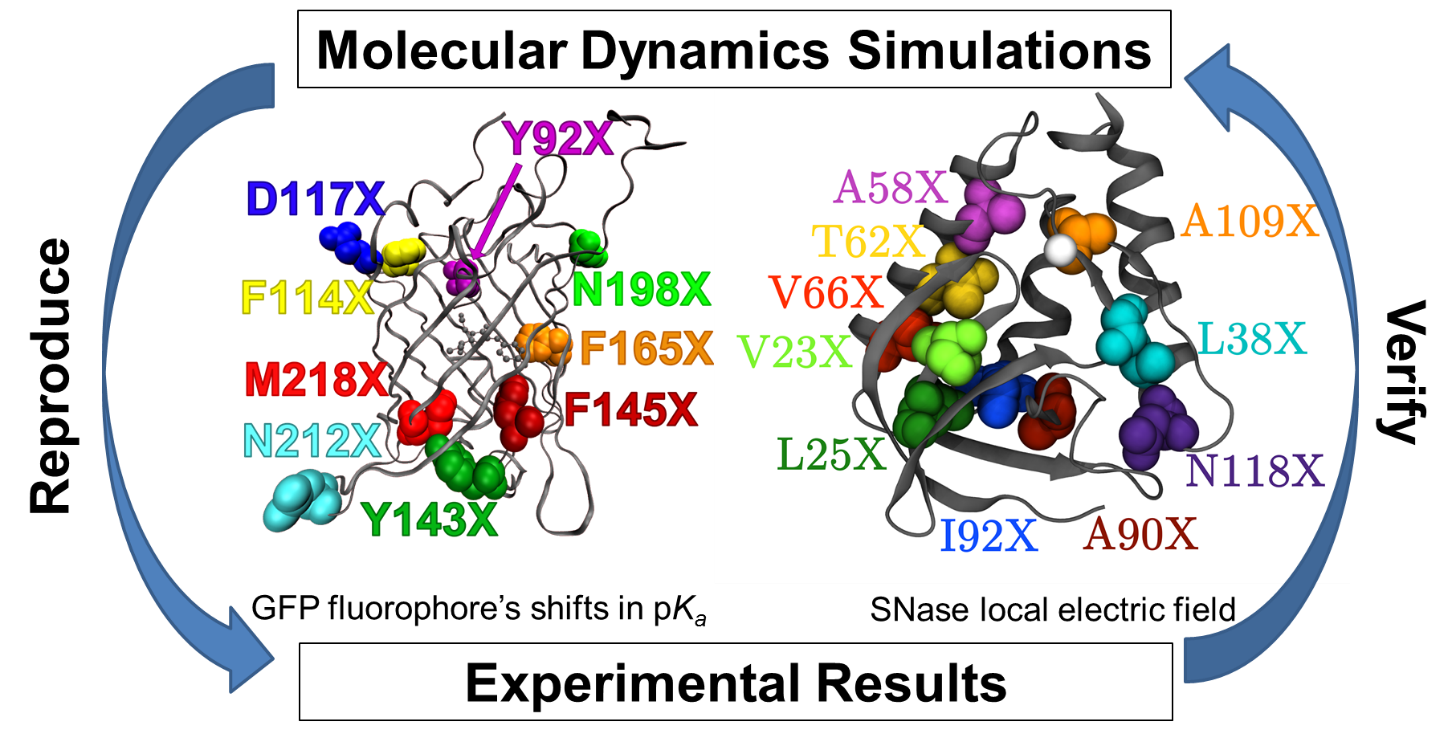
Molecular Dynamics Simulations Learn how md simulations capture the behavior of biomolecules in full atomic detail and at very fine temporal resolution. see how md simulations are used in molecular biology and drug discovery, especially in neuroscience, and how they complement experimental structural biology techniques. Molecular dynamics (md) is a method for analyzing the physical movements of atoms and molecules by numerically solving newton's equations of motion. md is applied in chemical physics, materials science, and biophysics to study the properties and dynamics of complex systems.
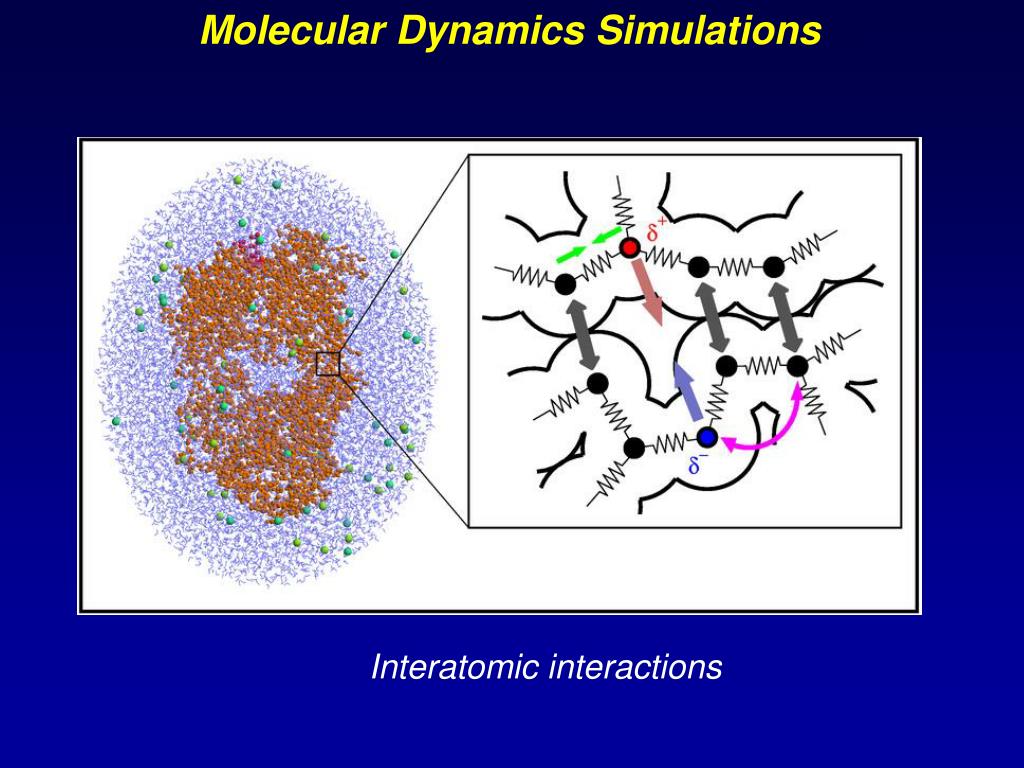
Ppt Molecular Dynamics Simulations Powerpoint Presentation Free Molecular dynamics (md) simulations have led to great advances in many scientific disciplines, such as chemical physics, materials science, and biophysics. This study comprehensively reviews the historical development, fundamental theories, and key techniques of md simulations, including all atom simulations, coarse grained simulations, and hybrid quantum mechanics molecular mechanics (qm mm) simulations. Driven by remarkable advances in computational power in recent years, molecular dynamics (md) simulations have become an indispensable tool in the research and development of materials, chemistry, drug discovery, and biosciences. Molecular dynamics is a method that uses newton’s equations of motion to computationally simulate the time evolution of a set of interacting atoms. such techniques are dependent on a.
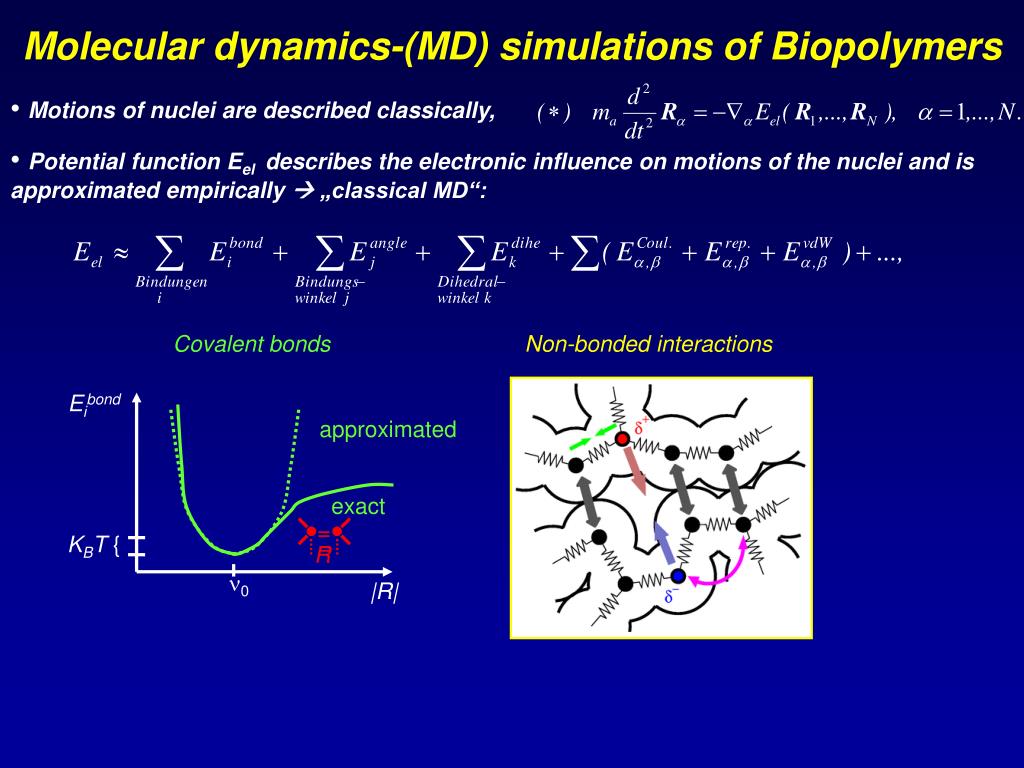
Ppt Molecular Dynamics Simulations Powerpoint Presentation Free Driven by remarkable advances in computational power in recent years, molecular dynamics (md) simulations have become an indispensable tool in the research and development of materials, chemistry, drug discovery, and biosciences. Molecular dynamics is a method that uses newton’s equations of motion to computationally simulate the time evolution of a set of interacting atoms. such techniques are dependent on a. Although md techniques are applied to a variety of biomolecules including dna, rna, proteins, and their assemblies, this review focuses specifically on the role of md in elucidating protein behavior and their interactions with inhibitors across different disease contexts. Molecular dynamics is defined as a simulation technique that generates the dynamic evolution of atoms in a molecular system over time by calculating atomic positions, velocities, and accelerations using newton's laws of motion. Learn the basic idea, equations, properties, applications, and limitations of molecular dynamics (md) simulations. md is a computational method that mimics the motion of atoms based on a potential energy function and newton's laws. Molecular dynamics (md) simulations have led to great advances in many scientific disciplines, such as chemical physics, materials science, and biophysics.
Comments are closed.