Models Linking Molecules To Organisms
Molecules And Organisms Vfx Unreal Engine Asset Assetsue These models then represent a significant first step toward a broader objective: to develop theoretical and computational models that describe how the physical interactions of molecules lead to complex biological phenotypes. Through the utilization of computational models, researchers are able to investigate the intricate dynamics of gene protein interactions and the involvement of other molecules, thereby facilitating a more profound comprehension of cellular processes and their regulatory mechanisms.

Molecules To Organisms Ngss Irvin schultz (battelle pacific nw national lab, marine sciences lab) gave a talk entitled "models linking molecules to organisms," at the predictive models. In this work, we aim at modeling and integrating several cellular processes into a single biochemical network, hence providing a more homogeneous framework to model whole organisms. Here, we present examples of foundational models developed at the organismal and ecological scale (lotka–volterra competition models and hardy–wienberg population genetics models) that have been successfully applied across biological scales to define, describe, and predict biological outcomes. This study presents a multiscale model of e. coli integrating metabolism and macromolecular synthesis, linking cellular physiology to codon usage evolution. it provides insights into the interplay between metabolic and translational processes.

Molecules To Organisms Ngss Here, we present examples of foundational models developed at the organismal and ecological scale (lotka–volterra competition models and hardy–wienberg population genetics models) that have been successfully applied across biological scales to define, describe, and predict biological outcomes. This study presents a multiscale model of e. coli integrating metabolism and macromolecular synthesis, linking cellular physiology to codon usage evolution. it provides insights into the interplay between metabolic and translational processes. In this context, complex systems and related interdisciplinary approaches of biology are expected to help. the aim of this collective paper is basically to provide a starting point for further. We describe how to build a discrete dynamic (boolean) model of a biological system from experimental data, and how to use the model to provide insights into emergent phenomena and make useful predictions. Models such as the “preferential attachment” model by barabási and albert (1999) and the evolving modularity model by valverde and solé (2007) have helped to understand how modular architectures can emerge in biological networks. As a complementary approach, genome scale metabolic models present a tractable way to integrate the effects of these additional mechanisms into predictions of community structure (31, 32). genome scale models are mathematical representations of the metabolic capabilities of individual organisms.

Unit 1 Molecules To Organisms Science With Mr Stiles In this context, complex systems and related interdisciplinary approaches of biology are expected to help. the aim of this collective paper is basically to provide a starting point for further. We describe how to build a discrete dynamic (boolean) model of a biological system from experimental data, and how to use the model to provide insights into emergent phenomena and make useful predictions. Models such as the “preferential attachment” model by barabási and albert (1999) and the evolving modularity model by valverde and solé (2007) have helped to understand how modular architectures can emerge in biological networks. As a complementary approach, genome scale metabolic models present a tractable way to integrate the effects of these additional mechanisms into predictions of community structure (31, 32). genome scale models are mathematical representations of the metabolic capabilities of individual organisms.
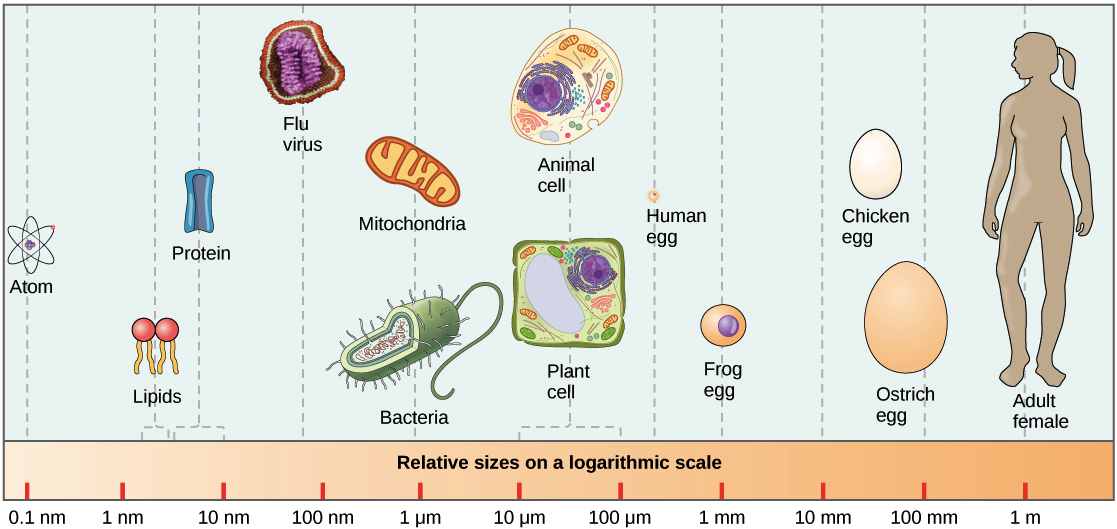
Molecules To Organisms Exploring Science Models such as the “preferential attachment” model by barabási and albert (1999) and the evolving modularity model by valverde and solé (2007) have helped to understand how modular architectures can emerge in biological networks. As a complementary approach, genome scale metabolic models present a tractable way to integrate the effects of these additional mechanisms into predictions of community structure (31, 32). genome scale models are mathematical representations of the metabolic capabilities of individual organisms.
Comments are closed.