Mnova Physchem Starting Guide

Physchem Validated and quantitative structure property prediction now integrated in mnova!. This advanced plugin needs a specific license (but it does not need to be installed apart).

Starting Guide To Mnova Db Mestrelab Research Analytical Chemistry Mnova does not perform automatic baseline corrections, so these should be done manually. click on the arrow by and then ‘baseline correction’ to get an interactive gui (shortcut key ‘b’). a dark blue line will show how the chosen method will fit your baseline. This document provides a starting guide for using mnova 12, which includes tutorials for processing 1d and 2d nmr data, multiplet analysis, peak assignment, and reporting results. Installation and activation of mnova, and general setup* *you will need to have mnova suite (nmr, nmrpredict, mschrom, and elvis) licenses for this tutorial. for the advanced tutorial, you will also need mnova qnmr and reaction monitoring licenses. Right click on the molecule structure and select ‘predict spectrum’ (1 h, 13 c, 31 p, 19 f, 15 n, 17 o, 29 si). then you will obtain the desired predicted spectrum as shown below: the spectrum can be analyzed as a real one (e.g. it can be integrated, peak picking, etc).

Starting Guide To Mnova Db Mestrelab Resources Installation and activation of mnova, and general setup* *you will need to have mnova suite (nmr, nmrpredict, mschrom, and elvis) licenses for this tutorial. for the advanced tutorial, you will also need mnova qnmr and reaction monitoring licenses. Right click on the molecule structure and select ‘predict spectrum’ (1 h, 13 c, 31 p, 19 f, 15 n, 17 o, 29 si). then you will obtain the desired predicted spectrum as shown below: the spectrum can be analyzed as a real one (e.g. it can be integrated, peak picking, etc). By default, mnova does the following automatically: picks peaks using gsd (if no peaks were picked) and classify their types (compound, solvent, impurity peaks etc.). The manual threshold option (shortcut k) allows you to select groups of peaks with different thresholds. the peak by peak option (ctrl k) is useful if you have shoulder peaks or ‘hidden peaks’ that were not selected in automatic peak picking. Mnova can analyze and report multiplets in terms of chemical shift, splitting pattern, j coupling(s), and # of protons. click analysis, multiplet analysis, automatic, then mnova will automatically select and analyze all peaks in the spectrum. Want to do something not covered in this mnova quick start guide? look here for a wealth of short and specific training materials!.
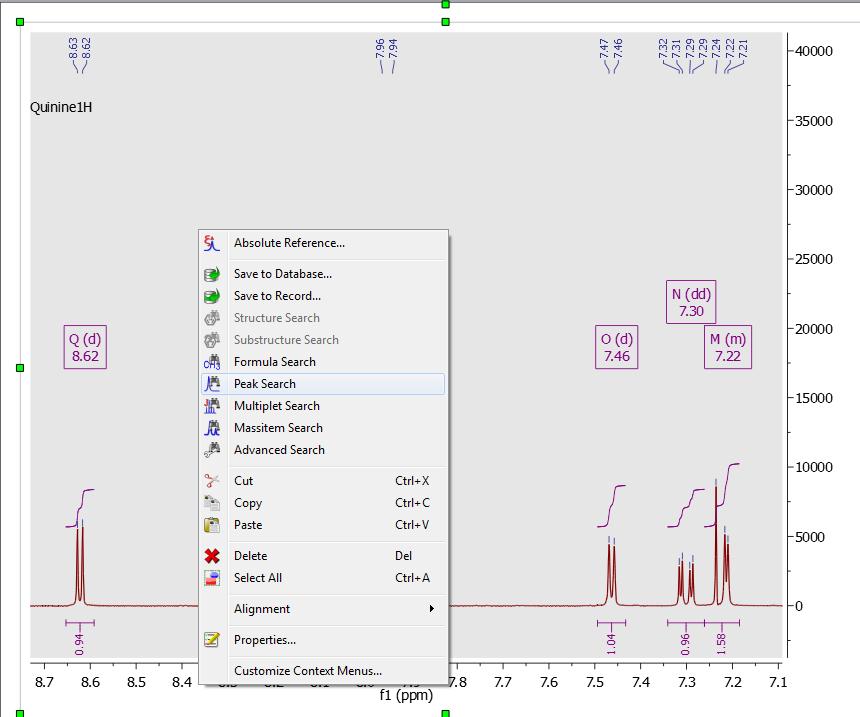
Starting Guide To Mnova Db Mestrelab Resources By default, mnova does the following automatically: picks peaks using gsd (if no peaks were picked) and classify their types (compound, solvent, impurity peaks etc.). The manual threshold option (shortcut k) allows you to select groups of peaks with different thresholds. the peak by peak option (ctrl k) is useful if you have shoulder peaks or ‘hidden peaks’ that were not selected in automatic peak picking. Mnova can analyze and report multiplets in terms of chemical shift, splitting pattern, j coupling(s), and # of protons. click analysis, multiplet analysis, automatic, then mnova will automatically select and analyze all peaks in the spectrum. Want to do something not covered in this mnova quick start guide? look here for a wealth of short and specific training materials!.

Mnova Iupac Name Starting Guide Mestrelab Research Analytical Mnova can analyze and report multiplets in terms of chemical shift, splitting pattern, j coupling(s), and # of protons. click analysis, multiplet analysis, automatic, then mnova will automatically select and analyze all peaks in the spectrum. Want to do something not covered in this mnova quick start guide? look here for a wealth of short and specific training materials!.

Starting Guide To Mnova Verify Mestrelab Resources
Comments are closed.