Mia Alejandro Reyes Yifeng Qi 3d Modelling Of Chromosomes In Cancer Nuclei
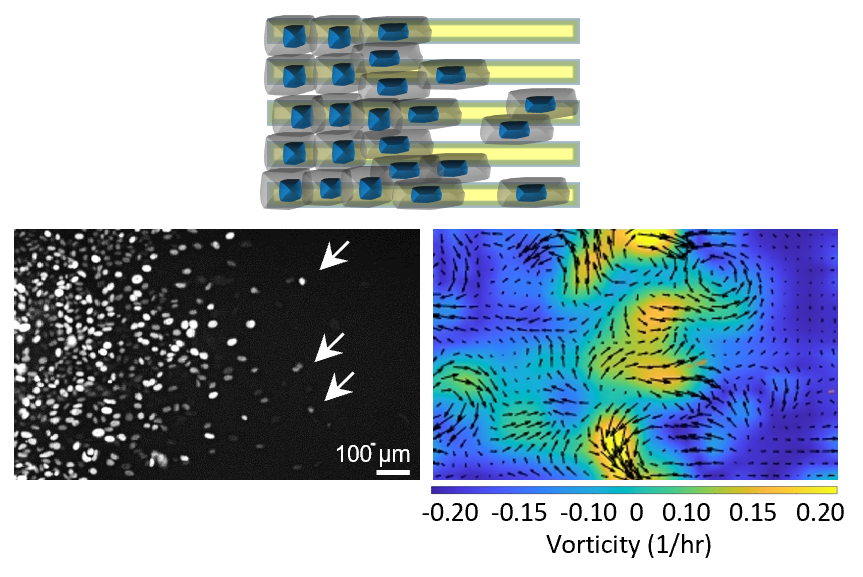
Research Chia Yi Su Lab Abstract: 3d maps of the human genome have revealed a hierarchical organization of dna within the nucleus. this organization, including chromatin loops, topologically associated domains (tads) and active and inactive compartments, has a strong association with transcription. We show an example where we have investigated tumor specific aberrations to 3d genome structure showing widespread disruption of heterochromatin organization in colon cancer.

Molecular Structure Of Cancer Cells Magnified Generated By Ai 25117172 Se johnstone, a reyes, y qi, c adriaens, e hegazi, k pelka, jh chen, l xie, p dong, x chen, ths hsieh, s banala, m de marzio, bp english, l xie, p dong, y qi, ths hsieh, bp. Three dimensional (3d) organization of the human genome plays an essential role in all dna templated processes, including gene transcription, gene regulation, and dna replication. Data driven polymer model for mechanistic exploration of diploid genome organization yifeng qi, alejandro reyes, sarah e johnstone, martin j aryee, bradley e bernstein, bin zhang. Determining how chromosomes are positioned and folded within the nucleus is critical to understanding the role of chromatin topology in gene regulation. several methods are available for.
Molecular Structure Of Cancer Cell Under Magnification Generated By Ai Data driven polymer model for mechanistic exploration of diploid genome organization yifeng qi, alejandro reyes, sarah e johnstone, martin j aryee, bradley e bernstein, bin zhang. Determining how chromosomes are positioned and folded within the nucleus is critical to understanding the role of chromatin topology in gene regulation. several methods are available for. Contributors: liangqi xie; peng dong; yifeng qi; tsung han s. hsieh; brian p. english; seolkyoung jung; xingqi chen; margherita de marzio; rafael casellas; howard y. chang et al. We introduce a predictive model to study cell type specific 3d chromatin folding. this model takes a sequence of chromatin states derived from genome wide histone modification profiles and a list of ctcf binding sites as input. Inter chromosomal interactions remains to be shown. on the other hand, imaging based techniques, though ideally suited for spatial measurements, are often low throughput and can face challenges for mechanistic explorations that require large data set collection for statistical significance. Here, we review a variety of large scale polymer models developed to investigate the structures and dynamics of chromosomes. different from the underlying modeling strategies, these approaches can be classified into data driven (‘top down’) and physics based (‘bottom up’) categories.
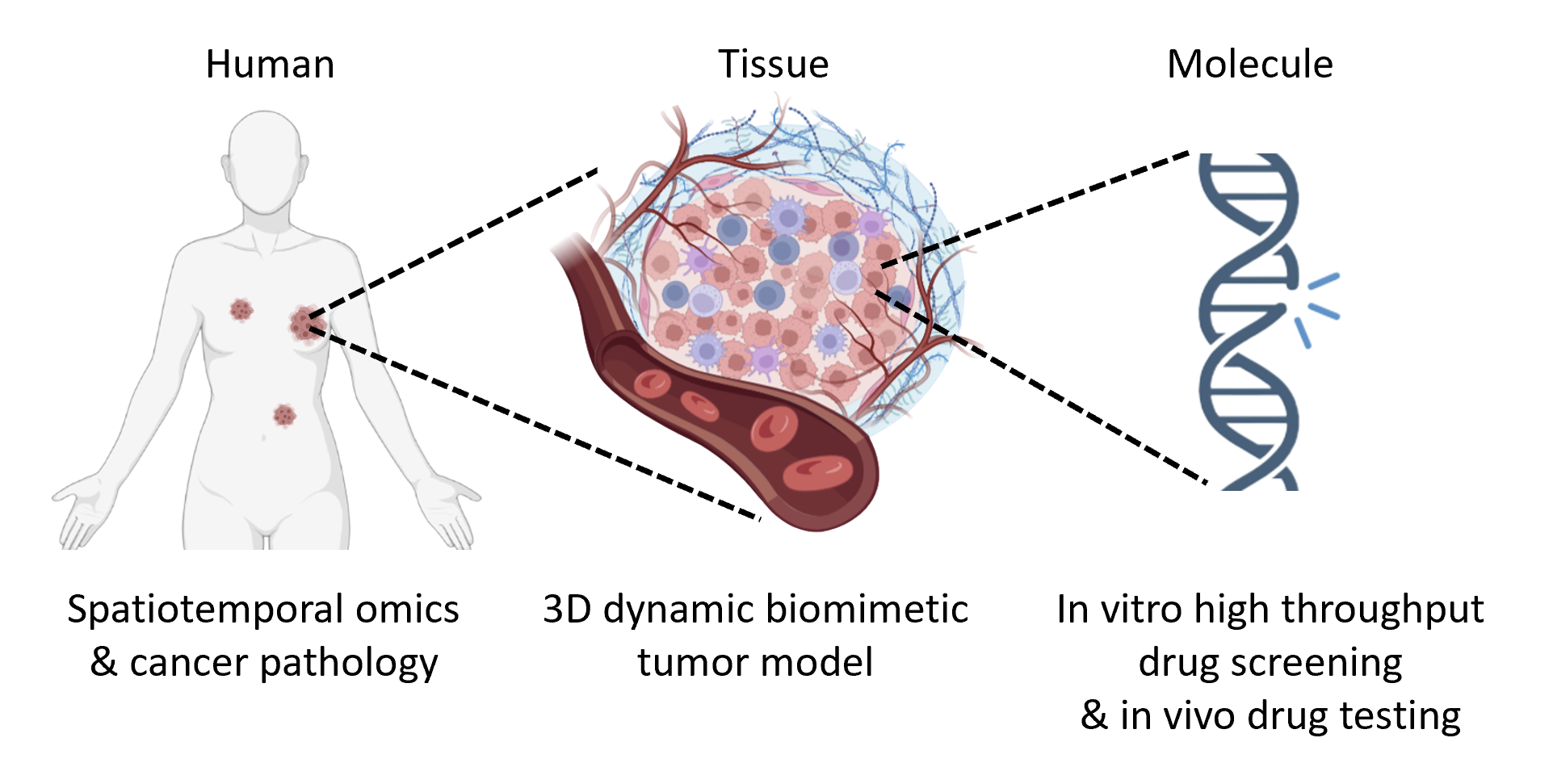
Chia Yi Su Lab Join Us To Fight Cancer Contributors: liangqi xie; peng dong; yifeng qi; tsung han s. hsieh; brian p. english; seolkyoung jung; xingqi chen; margherita de marzio; rafael casellas; howard y. chang et al. We introduce a predictive model to study cell type specific 3d chromatin folding. this model takes a sequence of chromatin states derived from genome wide histone modification profiles and a list of ctcf binding sites as input. Inter chromosomal interactions remains to be shown. on the other hand, imaging based techniques, though ideally suited for spatial measurements, are often low throughput and can face challenges for mechanistic explorations that require large data set collection for statistical significance. Here, we review a variety of large scale polymer models developed to investigate the structures and dynamics of chromosomes. different from the underlying modeling strategies, these approaches can be classified into data driven (‘top down’) and physics based (‘bottom up’) categories.
Comments are closed.