Imetalab
Imetalab Imetalab represents the whole framework, including the metalab desktop version, automatic imetareport, imetashiny apps for data visualization and analysis, metax for taxon function cross analysis. Imetalab is a cloud platform, which is developed by dr. daniel figeys research group at university of ottawa, for metaproteomics data analysis.

Imetalab Framework of the imetalab suite. users load raw files to the metalab desktop software to perform an automated metaproteomics database search. after the search, a series of result tables will be generated. Imetalab is a cloud based platform providing a user friendly web interface allowing general users to directly obtain peptide and taxa abundance information and protein function annotation from raw ms data. imetalab platform was originally released on september 13, 2017. If you would like to have a quick look at imetalab's data processing features without setting up your own session, please feel free to check out our demonstration session. With imetalab suite, we aim to maximize the accessibility of metaproteomic bioinformatics workflow to scientists with all levels of bioinformatics expertise in the field of microbiome research, as well as those in conventional proteomics systems biology.

Imetalab If you would like to have a quick look at imetalab's data processing features without setting up your own session, please feel free to check out our demonstration session. With imetalab suite, we aim to maximize the accessibility of metaproteomic bioinformatics workflow to scientists with all levels of bioinformatics expertise in the field of microbiome research, as well as those in conventional proteomics systems biology. To overcome these challenges, we have developed a set of tools named imetalab suite. imetalab suite includes the following components: (1) metalab desktop, an automated database search software that facilities proteins identification and quantitation from microbiomes; (2) the automated imetareport that allows users to quickly access database. A peptide focused functional enrichment workflow for gut microbiome metaproteomic studies figure oriented organization of code to plot figures plot a few particular figures hard to render elsewhere advanced visualization of the taxonomy output generated by metalab batch effect explore and correction for your dataset. We share the toolset with the microbiome research community. imetalab suite now has registered users from over 160 different institutions around the world. we aim to make imetalab suite a free and one stop toolset for metaproteomics, with increasing amounts of tools under active development. To overcome these challenges, we have developed a set of tools named imetalab suite. imetalab suite includes the following components: (1) metalab desktop, an automated database search software that facilities proteins identification and quantitation from microbiomes; (2) the automated imetareport that allows users to quickly access database.
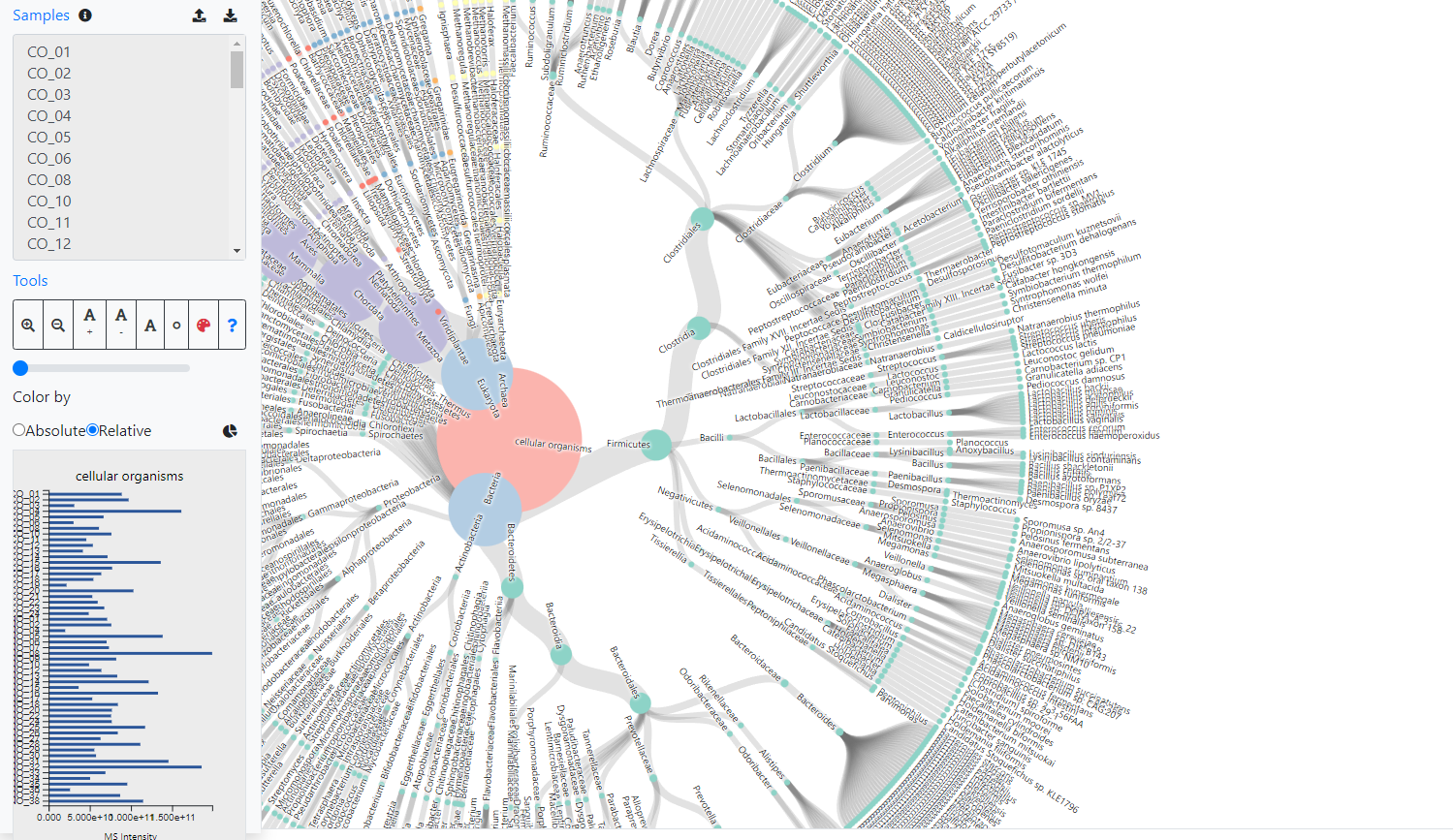
Imetalab To overcome these challenges, we have developed a set of tools named imetalab suite. imetalab suite includes the following components: (1) metalab desktop, an automated database search software that facilities proteins identification and quantitation from microbiomes; (2) the automated imetareport that allows users to quickly access database. A peptide focused functional enrichment workflow for gut microbiome metaproteomic studies figure oriented organization of code to plot figures plot a few particular figures hard to render elsewhere advanced visualization of the taxonomy output generated by metalab batch effect explore and correction for your dataset. We share the toolset with the microbiome research community. imetalab suite now has registered users from over 160 different institutions around the world. we aim to make imetalab suite a free and one stop toolset for metaproteomics, with increasing amounts of tools under active development. To overcome these challenges, we have developed a set of tools named imetalab suite. imetalab suite includes the following components: (1) metalab desktop, an automated database search software that facilities proteins identification and quantitation from microbiomes; (2) the automated imetareport that allows users to quickly access database.
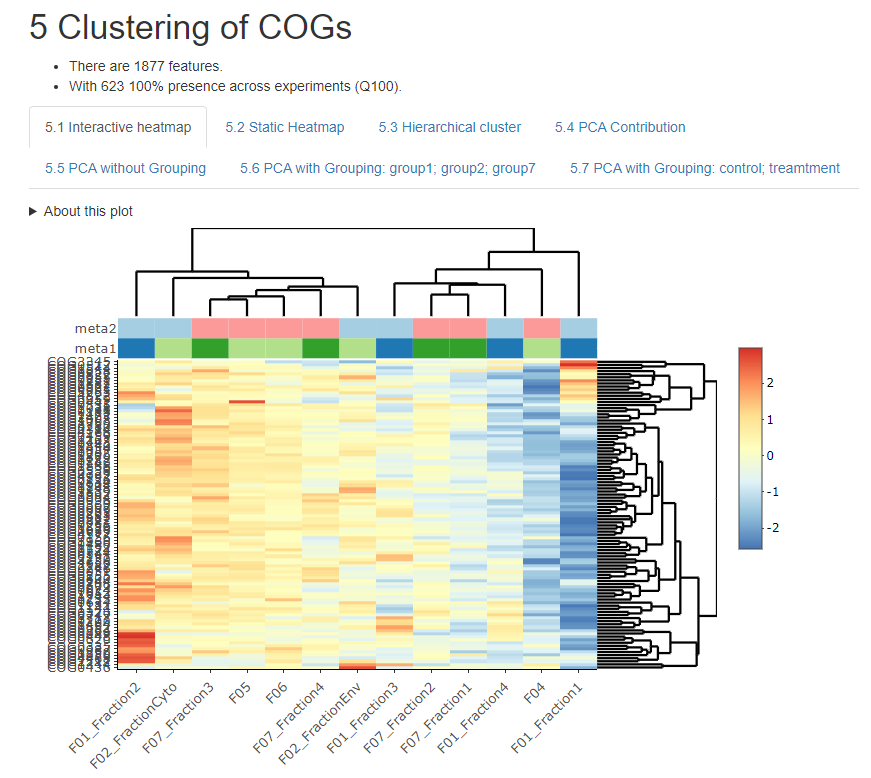
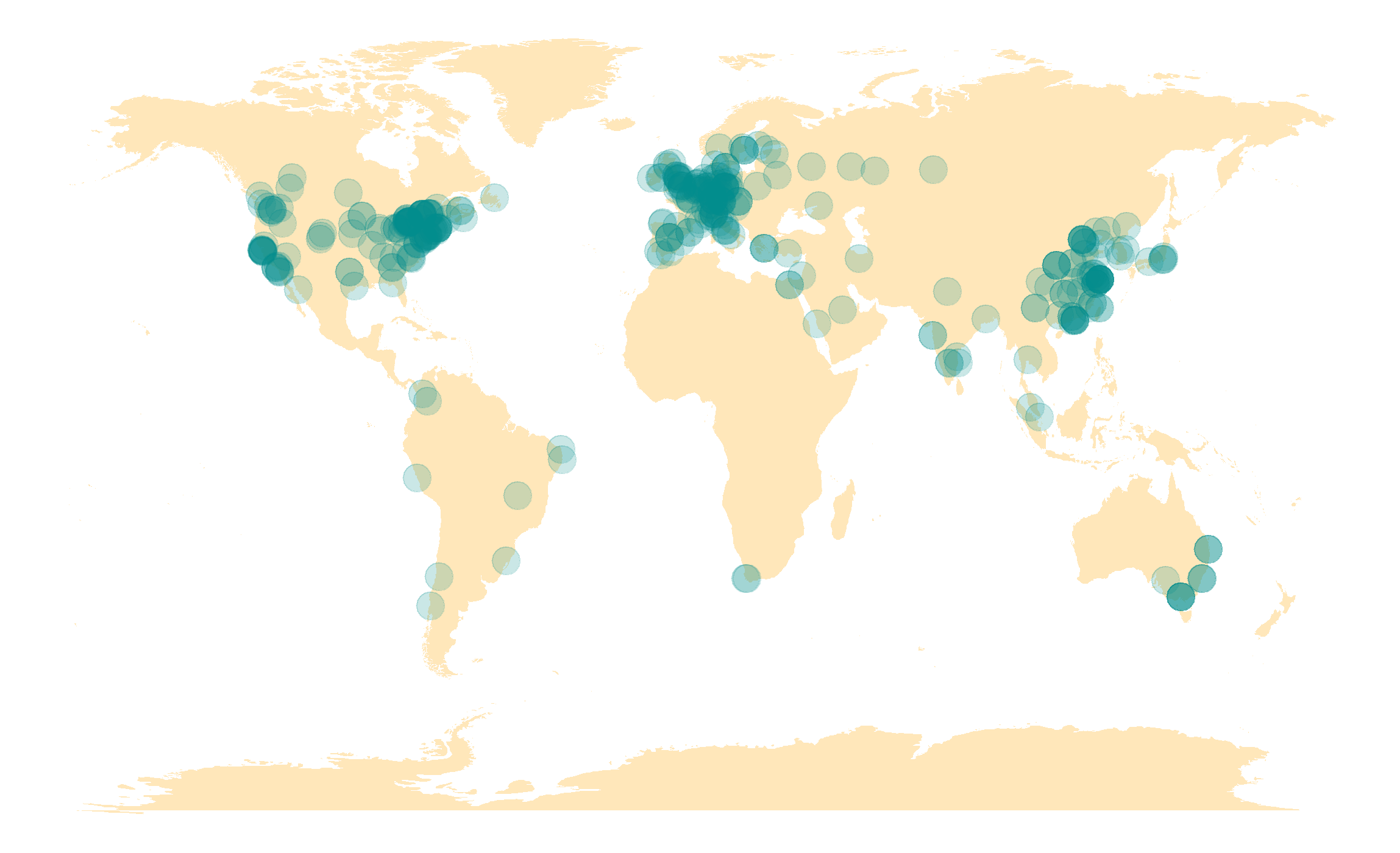
Imetalab We share the toolset with the microbiome research community. imetalab suite now has registered users from over 160 different institutions around the world. we aim to make imetalab suite a free and one stop toolset for metaproteomics, with increasing amounts of tools under active development. To overcome these challenges, we have developed a set of tools named imetalab suite. imetalab suite includes the following components: (1) metalab desktop, an automated database search software that facilities proteins identification and quantitation from microbiomes; (2) the automated imetareport that allows users to quickly access database.
Imetalab
Comments are closed.