Github Pnnl Compbio Neato
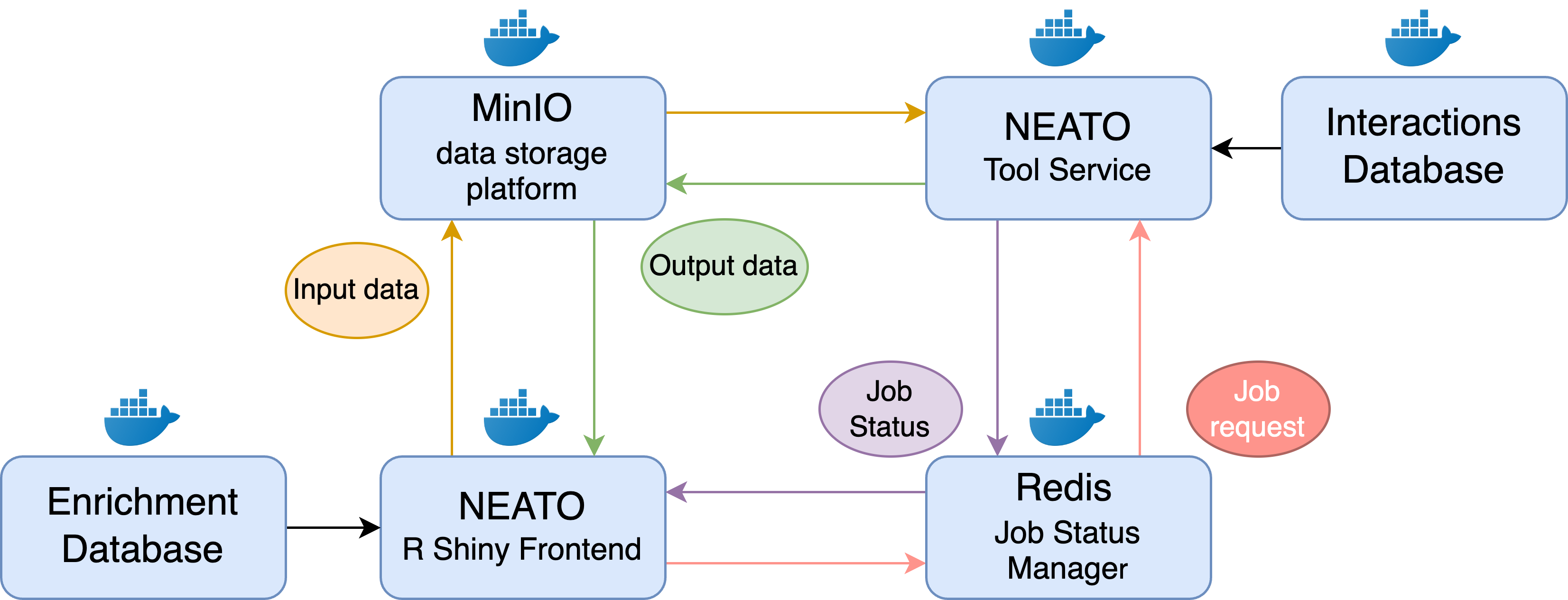
Github Pnnl Compbio Neato This data is used in combination with the interactions stored in the string database, which will be housed locally, to contruct a network using one of the provided algorithms hosted in the neato service contianer. Have a question about this project? by clicking “sign up for github”, you agree to our terms of service and privacy statement. we’ll occasionally send you account related emails. already on github? sign in to your account 5 open 3 closed 5 open 3 closed assignee.
Computational Biology Pacific Northwest National Laboratory Github Contribute to pnnl compbio neato development by creating an account on github. There aren’t any open pull requests. you could search all of github or try an advanced search. protip!. Contribute to pnnl compbio neato development by creating an account on github. Have a question about this project? sign up for a free github account to open an issue and contact its maintainers and the community. by clicking “sign up for github”, you agree to our terms of service and privacy statement. we’ll occasionally send you account related emails. already on github? sign in to your account.
Github Pnnl Compbio Boltzmannmfx Boltzmannmfx Is A Biological Contribute to pnnl compbio neato development by creating an account on github. Have a question about this project? sign up for a free github account to open an issue and contact its maintainers and the community. by clicking “sign up for github”, you agree to our terms of service and privacy statement. we’ll occasionally send you account related emails. already on github? sign in to your account. To facilitate the development as well as improve the reproducibility and reusability of multi omic deconvolution algorithms, we introduce decomprolute, a common workflow language framework that leverages containerization to compare tumor deconvolution algorithms across multiomic data sets. Github is where people build software. more than 150 million people use github to discover, fork, and contribute to over 420 million projects. Release for feng et al, 2024 manuscript. Core pnnl research capabilities captured resolve into five research areas: 1) chemical and materials sciences, 2) computational and mathematical sciences, 3) earth and biological sciences, 4) engineering, and 5) user facilities and advanced instrumentation.

Github Pnnl Compbio Ion Mob Ms Workflow For Analyzing Ion Mobility To facilitate the development as well as improve the reproducibility and reusability of multi omic deconvolution algorithms, we introduce decomprolute, a common workflow language framework that leverages containerization to compare tumor deconvolution algorithms across multiomic data sets. Github is where people build software. more than 150 million people use github to discover, fork, and contribute to over 420 million projects. Release for feng et al, 2024 manuscript. Core pnnl research capabilities captured resolve into five research areas: 1) chemical and materials sciences, 2) computational and mathematical sciences, 3) earth and biological sciences, 4) engineering, and 5) user facilities and advanced instrumentation.
Compbio Ai Github Release for feng et al, 2024 manuscript. Core pnnl research capabilities captured resolve into five research areas: 1) chemical and materials sciences, 2) computational and mathematical sciences, 3) earth and biological sciences, 4) engineering, and 5) user facilities and advanced instrumentation.
Yale Compbio Github
Comments are closed.