Github Micom Dev Micom Python Package To Study Microbial Communities
Github Micom Dev Micom Python Package To Study Microbial Communities Micom is a python package for metabolic modeling of microbial communities currently developed in the diener lab at the medical university of graz. Micom is a python package for metabolic modeling of microbial communities developed in the gibbons lab at the institute for systems biology and the human systems biology group of prof. osbaldo resendis antonio at the national institute of genomic medicine mexico.
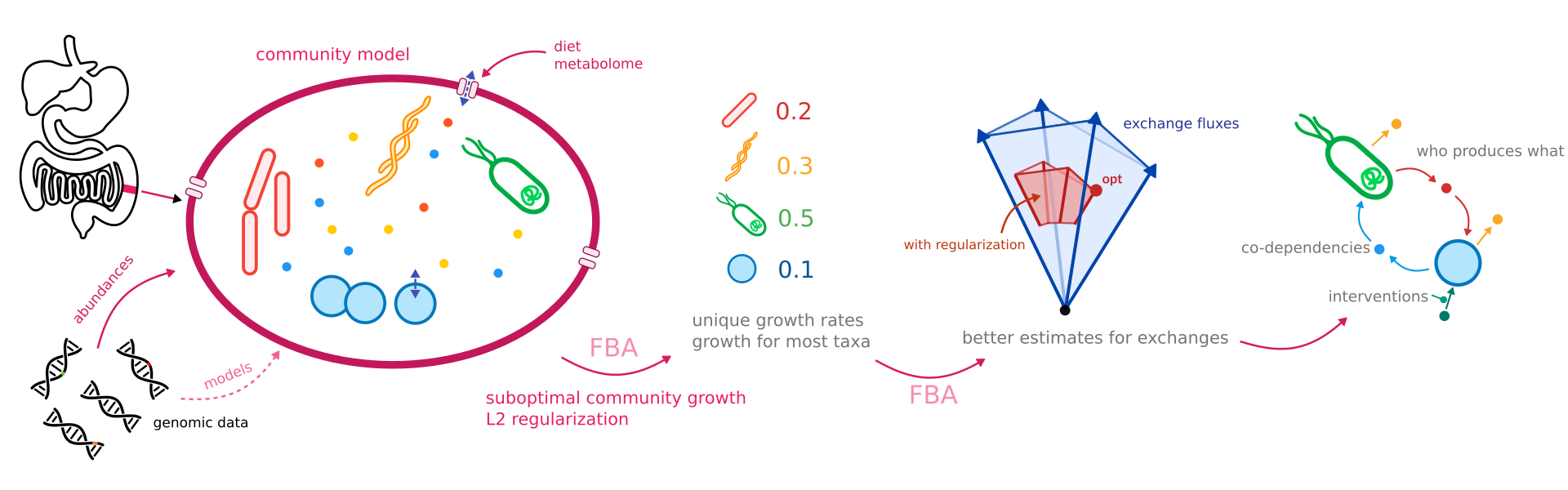
How Does Micom Model Communities Version 0 37 1 Micom is a python package for metabolic modeling of microbial communities currently developed in the diener lab at the medical university of graz. Workflows and input data for the construction of the standard micom model databases. environmental growth media for micom. materials and code for the micom manuscript. Micom is a python package for metabolic modeling of microbial communities currently developed in the diener lab at the medical university of graz. Python package to study microbial communities using metabolic modeling. releases · micom dev micom.
Micom Dev Github Micom is a python package for metabolic modeling of microbial communities currently developed in the diener lab at the medical university of graz. Python package to study microbial communities using metabolic modeling. releases · micom dev micom. Python package to study microbial communities using metabolic modeling. micom micom community.py at main · micom dev micom. Micom is a python package for metabolic modeling of microbial communities currently developed in the gibbons lab at the institute for systems biology and the human systems biology group of prof. osbaldo resendis antonio at the national institute of genomic medicine mexico. To make this tutorial as accessible as possible we will use an open source hybrid solver developed for micom. this solvr is accurate but may be slower than commercial solvers. To help with this micom provides a workflow that can complete any predefined growth medium with the minimal additional substrates to allow growth for all taxa in the database.
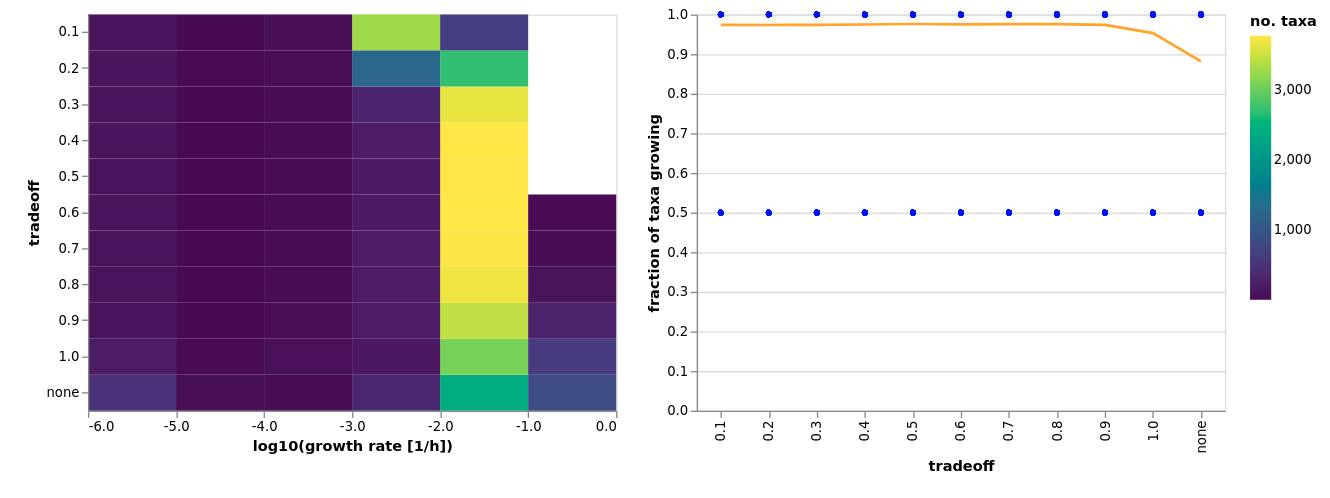
Basic Questions On Metabolic Modelling Of Microbial Communities Micom Python package to study microbial communities using metabolic modeling. micom micom community.py at main · micom dev micom. Micom is a python package for metabolic modeling of microbial communities currently developed in the gibbons lab at the institute for systems biology and the human systems biology group of prof. osbaldo resendis antonio at the national institute of genomic medicine mexico. To make this tutorial as accessible as possible we will use an open source hybrid solver developed for micom. this solvr is accurate but may be slower than commercial solvers. To help with this micom provides a workflow that can complete any predefined growth medium with the minimal additional substrates to allow growth for all taxa in the database.
Q2 Micom A Qiime Plugin For Micom To make this tutorial as accessible as possible we will use an open source hybrid solver developed for micom. this solvr is accurate but may be slower than commercial solvers. To help with this micom provides a workflow that can complete any predefined growth medium with the minimal additional substrates to allow growth for all taxa in the database.
Github Wisang Micom
Comments are closed.