Github Bioanalyticresource Eplant Plant Efp A Data Visualization
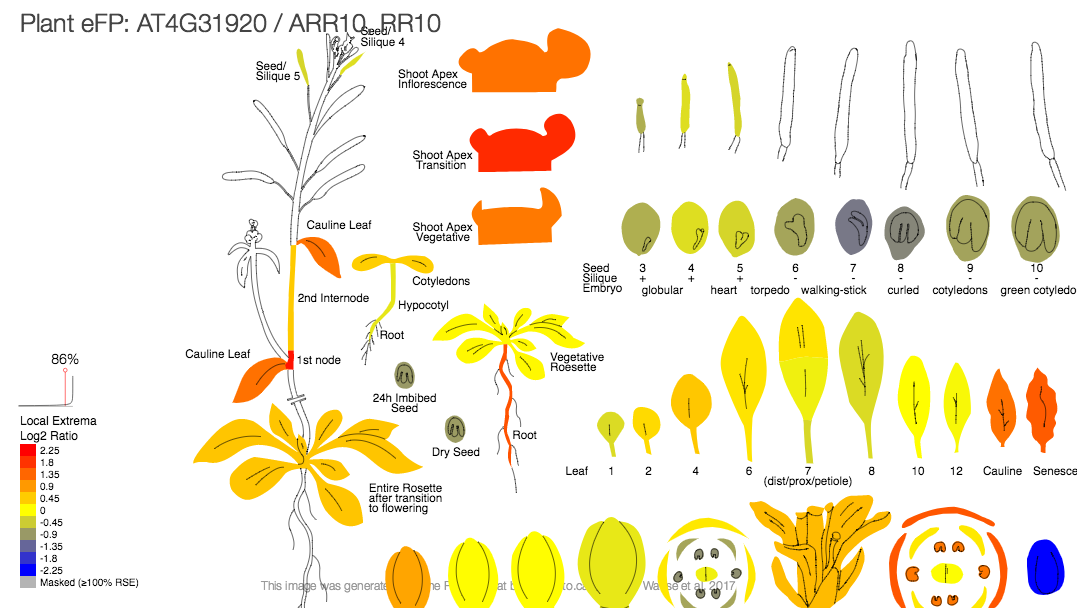
Eplant A Comprehensive Data Visualization Tool For Plant Biologists The eplant plant efp is a tissue expression api provided by the bio analytic resource for plant biology (bar) from the university of toronto. this tool will provide visualized tissue expression data corresponding for arabidopsis thaliana with one of the listed compendiums. An rna seq data exploration tool that shows read map coverage of a gene of interest along with a coloured "electronic fluorescent pictographic" (efp) based on its rpkm expression level.
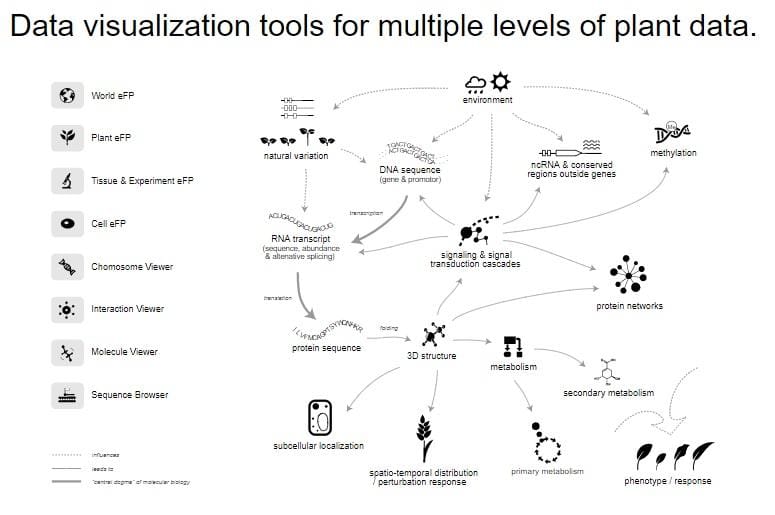
The Graphical User Interface That Greets You On Eplant S Home Page A data visualization tool to display tissue expression data for arabidopsis thaliana bioanalyticresource eplant plant efp. The eplant plant efp is a tissue expression api provided by the bio analytic resource for plant biology (bar) from the university of toronto. this tool will provide visualized tissue expression data corresponding for arabidopsis thaliana with one of the listed compendiums. Eplant is a gene centric visualization tool for plant genomes bioanalyticresource eplant. An rna seq data exploration tool that shows read map coverage of a gene of interest along with a coloured "electronic fluorescent pictographic" (efp) based on its rpkm expression level.

Subscribe Eplant is a gene centric visualization tool for plant genomes bioanalyticresource eplant. An rna seq data exploration tool that shows read map coverage of a gene of interest along with a coloured "electronic fluorescent pictographic" (efp) based on its rpkm expression level. The interactions viewer displays protein interactors for the currently selected gene product from our database of ~80k predicted and ~100k confirmed protein interactions, and 2.8m protein dna interactions. The gene info viewer view requires a selected gene. The ability to display gene expression profiles in plant efp viewers (fig. 1b) has proven to be a highly effective approach for visualization, communication, and education within the plant sciences. To fill the existing resource gap, we introduce ‘ggplantmap’, an open source r package with the goal of providing a simple and easy way to create efp like visualizations using the r language (fig. 2).
Plant Data Github The interactions viewer displays protein interactors for the currently selected gene product from our database of ~80k predicted and ~100k confirmed protein interactions, and 2.8m protein dna interactions. The gene info viewer view requires a selected gene. The ability to display gene expression profiles in plant efp viewers (fig. 1b) has proven to be a highly effective approach for visualization, communication, and education within the plant sciences. To fill the existing resource gap, we introduce ‘ggplantmap’, an open source r package with the goal of providing a simple and easy way to create efp like visualizations using the r language (fig. 2).
Comments are closed.