Gencode Human Genome Annotation Gff3 Kaggle

Gencode Human Genome Annotation Gff3 Kaggle Kaggle uses cookies from google to deliver and enhance the quality of its services and to analyze traffic. ok, got it. something went wrong and this page crashed! if the issue persists, it's likely a problem on our side. at kaggle static assets app.js?v=8a659a32a15e0ff4:1:2544755. Human release 46 (grch38.p14) statistics of this release more information about this assembly (including patches, scaffolds and haplotypes) go to grch37 version of this release gtf gff3 files fasta files metadata files.

Human Complete Genome Sequence Features Kaggle Get gencode annotation.sh is a script that downloads a specified gencode annotation in gff3 format and generates several useful bed files. we use it to generate the annotation files we use in our bioinformatic projects. Downloads a gff3 file for a specified species, release version, and annotation type from the gencode database. the file is saved to a user specified directory or the current working directory by default. The human gene annotations from the gencode project. (the 'comprehensive gene annotation' on the chr regions will do make sure to get the 'gff3' formatted files.). Download it from the gencode download page for human gene annotations you want the 'comprehensive gene annotation' file in gff3 format. or download the copy of the gencode file that i have placed in this folder.

Grch38 Human Genome Dna Kaggle The human gene annotations from the gencode project. (the 'comprehensive gene annotation' on the chr regions will do make sure to get the 'gff3' formatted files.). Download it from the gencode download page for human gene annotations you want the 'comprehensive gene annotation' file in gff3 format. or download the copy of the gencode file that i have placed in this folder. We aim to annotate all human and mouse protein coding genes, long non coding rnas (lncrnas), pseudogenes and small rnas (these are annotated by a computational pipeline linked to external databases and are not discussed further). Annotation transfer or "liftover" is an approach where, through a whole genome alignment between a high quality genome assembly (that has a similarly high quality annotation) and an unannotated target genome assembly, annotations are transferred to homologous intervals in the target genome. Gencode is a comprehensive annotation project that aims to provide high quality annotations of the human and mouse genomes. the project is part of the encode (encyclopedia of dna elements) scale up project, which seeks to identify all functional elements in the human genome. The sga files were generated from gff3 format with a custom shell script available from epd.expasy.org ftp mga hg38 gencode scripts in the latest version v47, all transcripts with "transcript support level=1" and "gene type=protein coding" were included.

Github Galikbence Genome Annotation Pipeline For Eukaryotic Genome We aim to annotate all human and mouse protein coding genes, long non coding rnas (lncrnas), pseudogenes and small rnas (these are annotated by a computational pipeline linked to external databases and are not discussed further). Annotation transfer or "liftover" is an approach where, through a whole genome alignment between a high quality genome assembly (that has a similarly high quality annotation) and an unannotated target genome assembly, annotations are transferred to homologous intervals in the target genome. Gencode is a comprehensive annotation project that aims to provide high quality annotations of the human and mouse genomes. the project is part of the encode (encyclopedia of dna elements) scale up project, which seeks to identify all functional elements in the human genome. The sga files were generated from gff3 format with a custom shell script available from epd.expasy.org ftp mga hg38 gencode scripts in the latest version v47, all transcripts with "transcript support level=1" and "gene type=protein coding" were included.
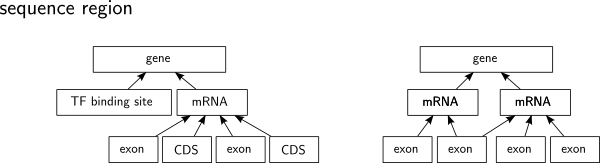
The Annotationsketch Module Gencode is a comprehensive annotation project that aims to provide high quality annotations of the human and mouse genomes. the project is part of the encode (encyclopedia of dna elements) scale up project, which seeks to identify all functional elements in the human genome. The sga files were generated from gff3 format with a custom shell script available from epd.expasy.org ftp mga hg38 gencode scripts in the latest version v47, all transcripts with "transcript support level=1" and "gene type=protein coding" were included.
Comments are closed.