Fair Edna
Fair Edna Adopting fair (findable, accessible, interoperable, reusable) data practices with edna data can transform monitoring of biodiversity and individual species, and support data driven biodiversity management across broad scales. Our work advances the goal to make edna data fair by identifying essential data components and formats, developing an edna specific, comprehensive metadata checklist that builds on existing biodiversity and molecular data standards, and providing data formatting guidelines for edna communities.
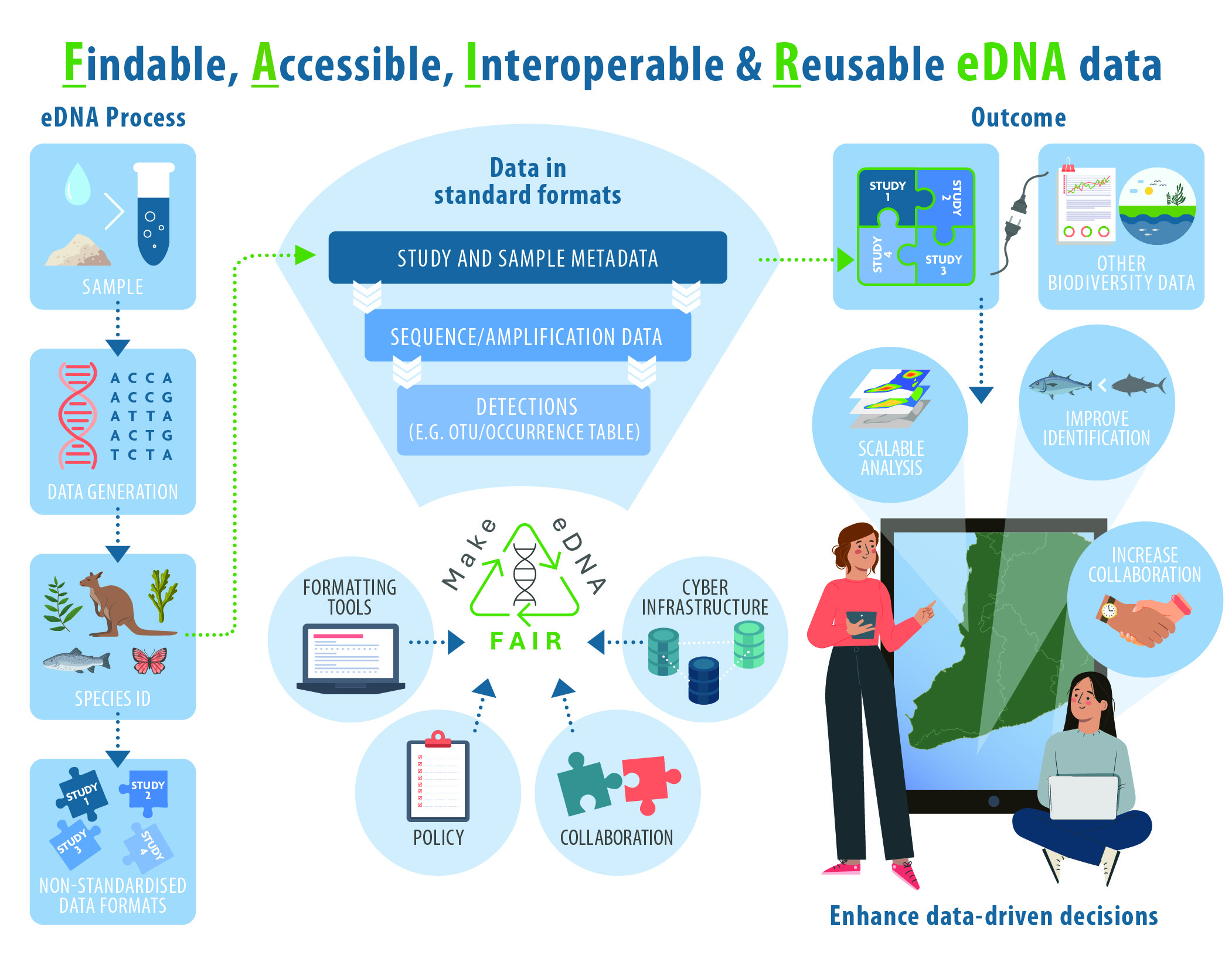
Fair Edna Collaborative development of the faire python package, consolidating and parsing existing and new python scripts into a unified package. the goal is to maintain feature parity with the r counterpart developed in the [faire r] ( github fair edna faire r) repository. Making edna data fair (findable, accessible, interoperable, re usable) has vast potential to improve how the biological environment is measured and how change is detected and understood. Environmental dna (edna) is a cutting edge scientific technique that involves collecting and analysing genetic material shed by organisms into their surrounding environment such as water and soil. Along with formatting guidelines, tools, templates, and example datasets, we introduce a standardized, ready‐to‐use approach for fair edna practices.

Edna Fashion Ednafashionoficial Threads Say More Environmental dna (edna) is a cutting edge scientific technique that involves collecting and analysing genetic material shed by organisms into their surrounding environment such as water and soil. Along with formatting guidelines, tools, templates, and example datasets, we introduce a standardized, ready‐to‐use approach for fair edna practices. We developed a fair edna (faire) metadata checklist, providing a comprehensive vocabulary for describing different data components and methodologies. visit the download tab to access the latest version of the checklist, the full template, and the previous versions with change logs. Adopting fair (findable, accessible, interoperable, and reusable) data practices with edna data can transform the monitoring of biodiversity and individual species and support data driven biodiversity management across broad scales. To address these challenges, we developed a comprehensive fair edna (faire) metadata checklist and formatting guidelines by synthesizing elements from existing standards such as dwc and mixs. these resources help edna practitioners structure their datasets according to fair principles. In this talk, we share our best practice guide for formatting and publishing edna data, developed by an international multidisciplinary working group comprising edna researchers, journal editors, and biodiversity and omics data scientists.
Fair Edna Github We developed a fair edna (faire) metadata checklist, providing a comprehensive vocabulary for describing different data components and methodologies. visit the download tab to access the latest version of the checklist, the full template, and the previous versions with change logs. Adopting fair (findable, accessible, interoperable, and reusable) data practices with edna data can transform the monitoring of biodiversity and individual species and support data driven biodiversity management across broad scales. To address these challenges, we developed a comprehensive fair edna (faire) metadata checklist and formatting guidelines by synthesizing elements from existing standards such as dwc and mixs. these resources help edna practitioners structure their datasets according to fair principles. In this talk, we share our best practice guide for formatting and publishing edna data, developed by an international multidisciplinary working group comprising edna researchers, journal editors, and biodiversity and omics data scientists.
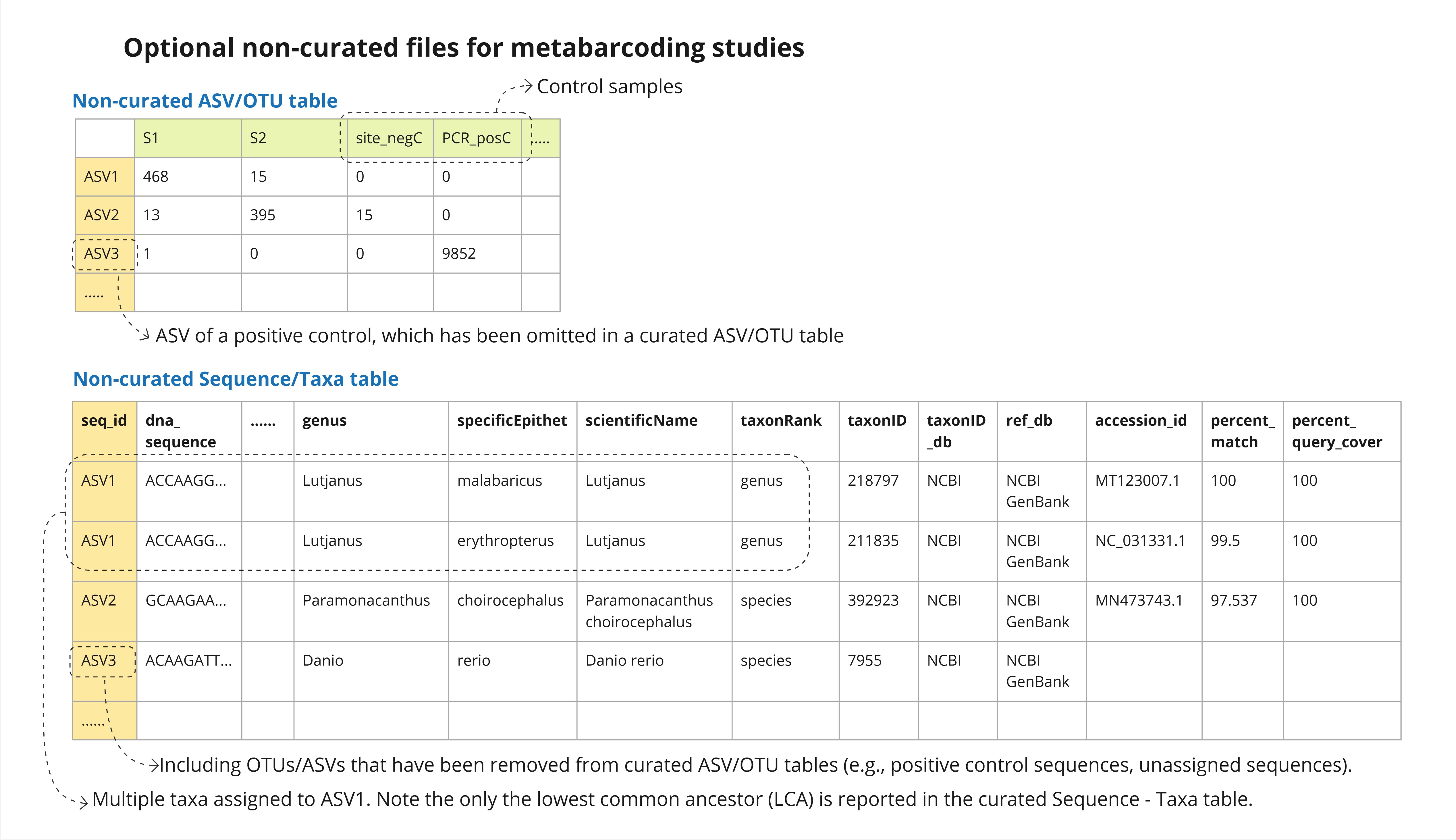
Guidelines Fair Edna To address these challenges, we developed a comprehensive fair edna (faire) metadata checklist and formatting guidelines by synthesizing elements from existing standards such as dwc and mixs. these resources help edna practitioners structure their datasets according to fair principles. In this talk, we share our best practice guide for formatting and publishing edna data, developed by an international multidisciplinary working group comprising edna researchers, journal editors, and biodiversity and omics data scientists.
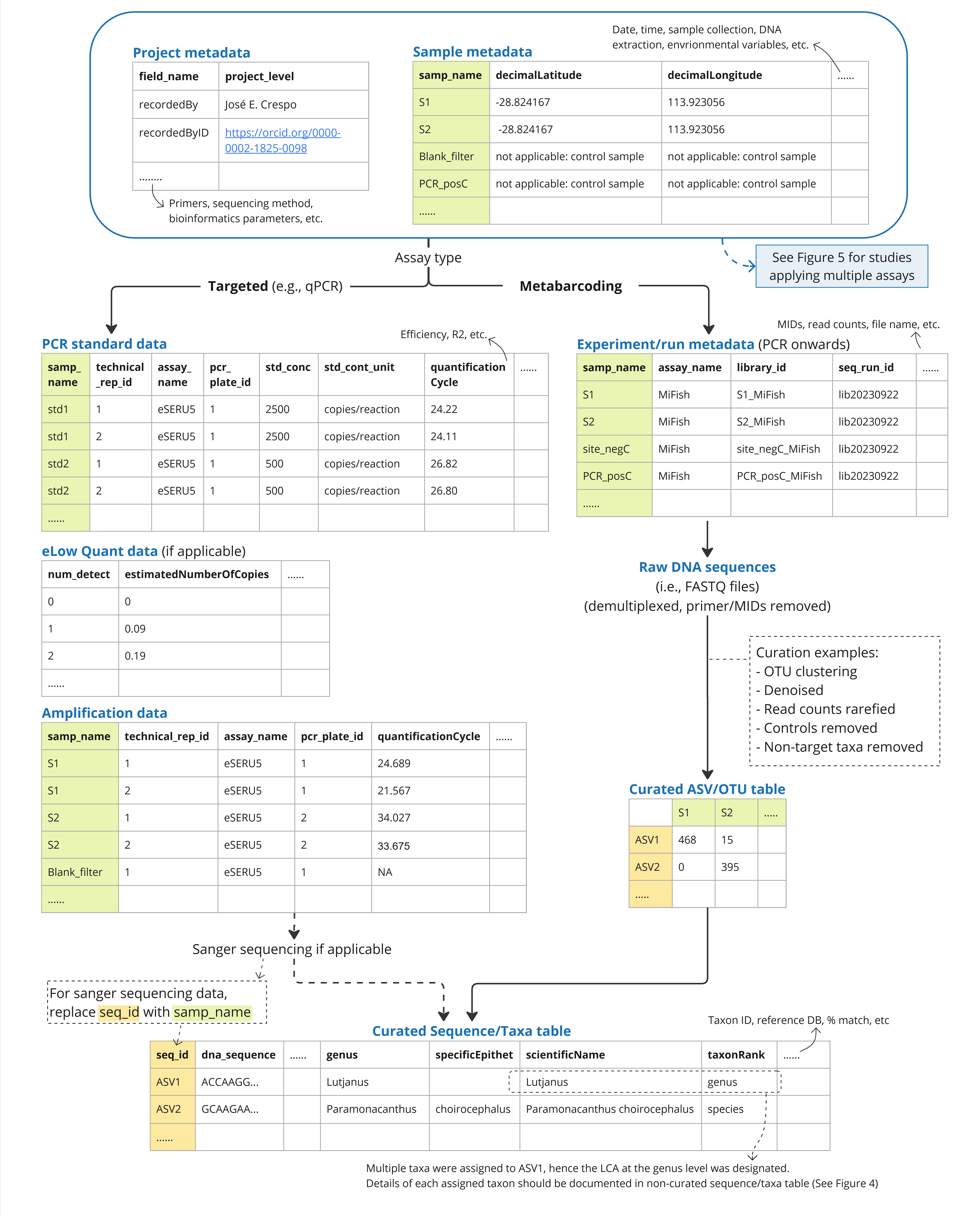
Guidelines Fair Edna
Comments are closed.