Evaluation Feature Allele
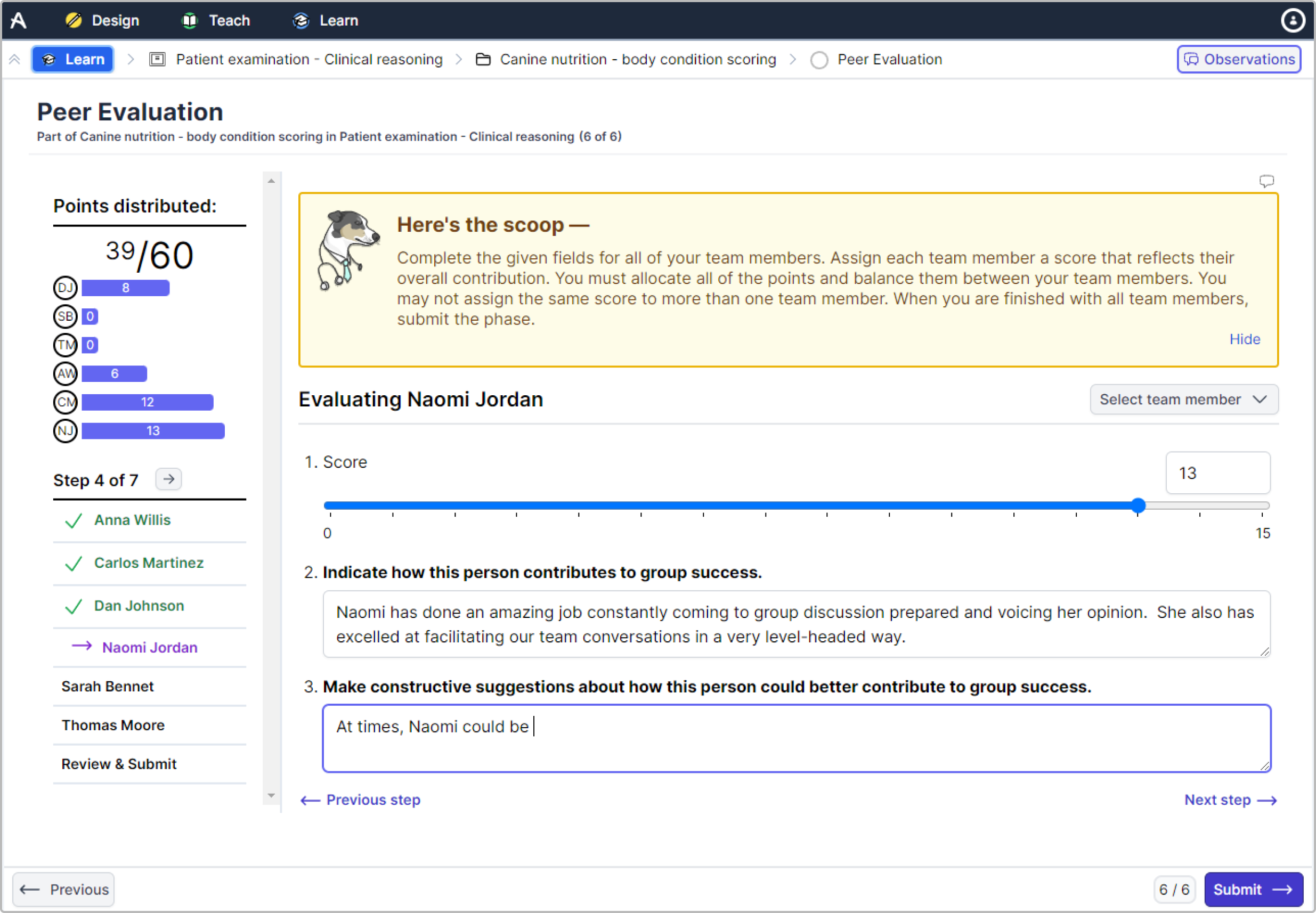
Evaluation Feature Allele You'll be sent a link to join and try allele at absolutely no cost. create engaging, real world case studies, pre work, application exercises, simulations, and clinical training. an instructional design and pedagogy software for teaching with high efficacy methodologies. Here, we benchmark feature selection methods for single cell rna sequencing integration using metrics beyond batch correction and preservation of biological variation to assess query mapping,.
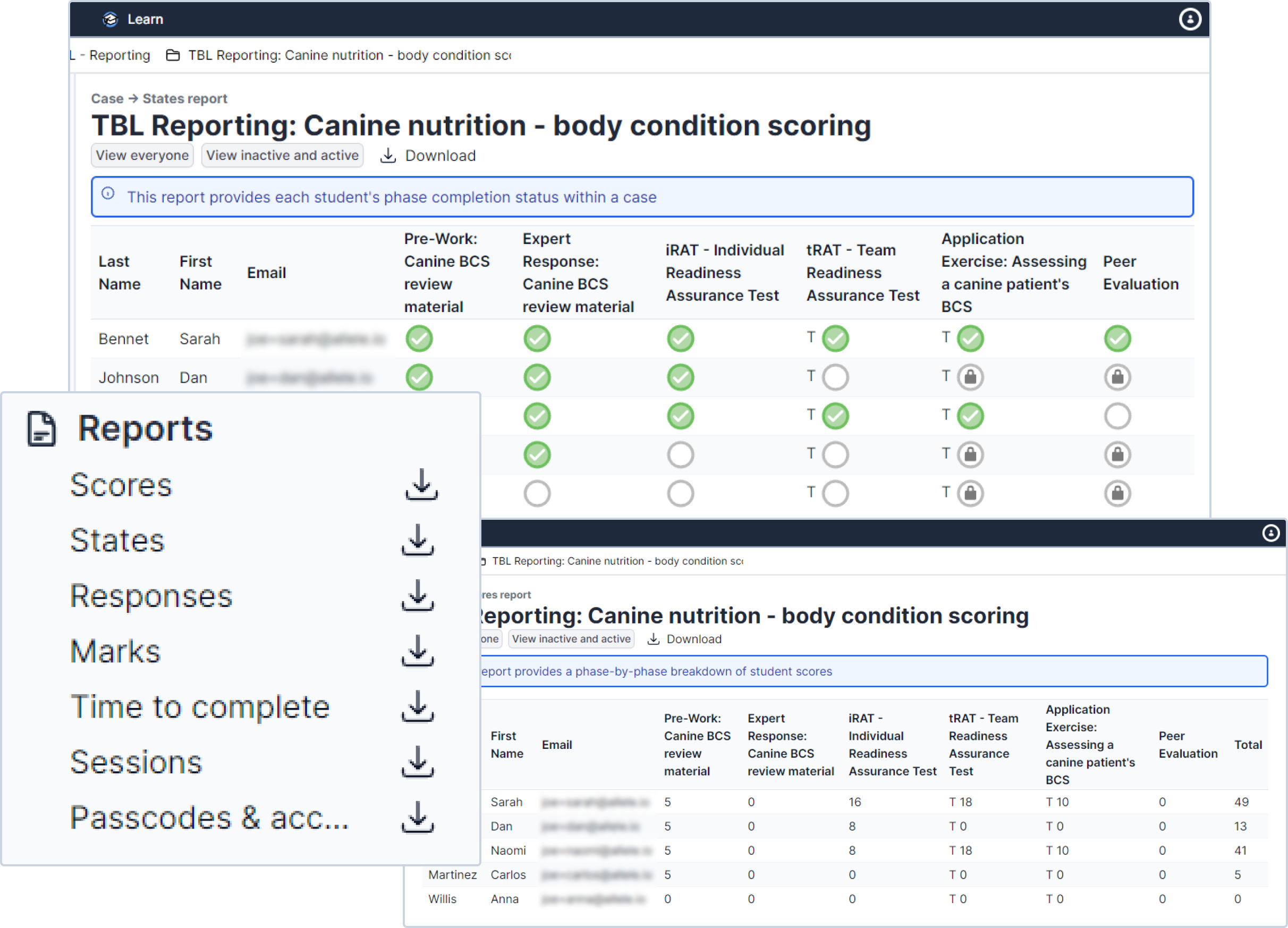
Reporting Feature Allele In this context, allele specific expression analysis serves as a powerful tool for understanding gene regulation, with significant functional and clinical implications. In this article, we provide a general overview of the different feature selection methods, their advantages, disadvantages, and use cases, focusing on the detection of relevant features (i.e., snps) for disease risk prediction. Each of the four panels details the percentage of alleles for which the nucleotide sequence of sets of gene features (gfs) is known at the hla a, b, c, or drb1 locus. This workflow adds new allele specific info level annotations to the vcfs and produces a final output with allele specific filters based on the vqsr snp and indel tranches.
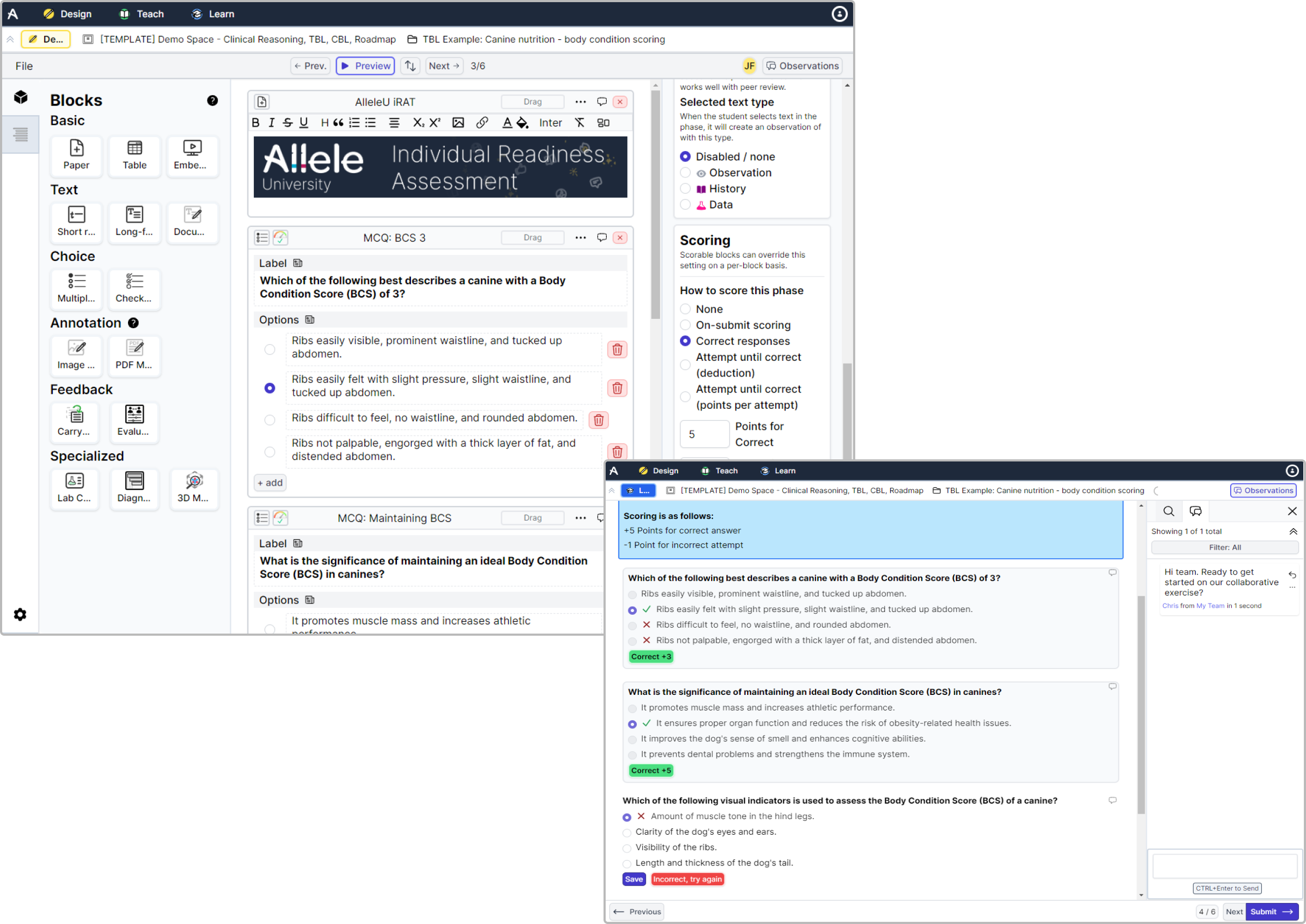
Scored Assessments Feature Allele Each of the four panels details the percentage of alleles for which the nucleotide sequence of sets of gene features (gfs) is known at the hla a, b, c, or drb1 locus. This workflow adds new allele specific info level annotations to the vcfs and produces a final output with allele specific filters based on the vqsr snp and indel tranches. This repository provides a framework for fine grained prediction of chromatin accessibility from dna sequence, using convnext v2 blocks as powerful feature extractors. The report provides summary statistics to evaluate results per sample and per locus (with the possibility to provide a tsv file with loci annotations to include on a table). For this purpose, we developed the maxx (mutation allelic expression extractor) software, which is highly effective at delineating the allelic expression of both single nucleotide variants and small insertions and deletions. In this article, we reviewed methods for detecting ase using both bulk rnaseq and single cell rnaseq data to provide deeper insights beyond the mere prediction of ase genes. fundamentally, ase detection methods are data driven and can be classified according to type of data used.
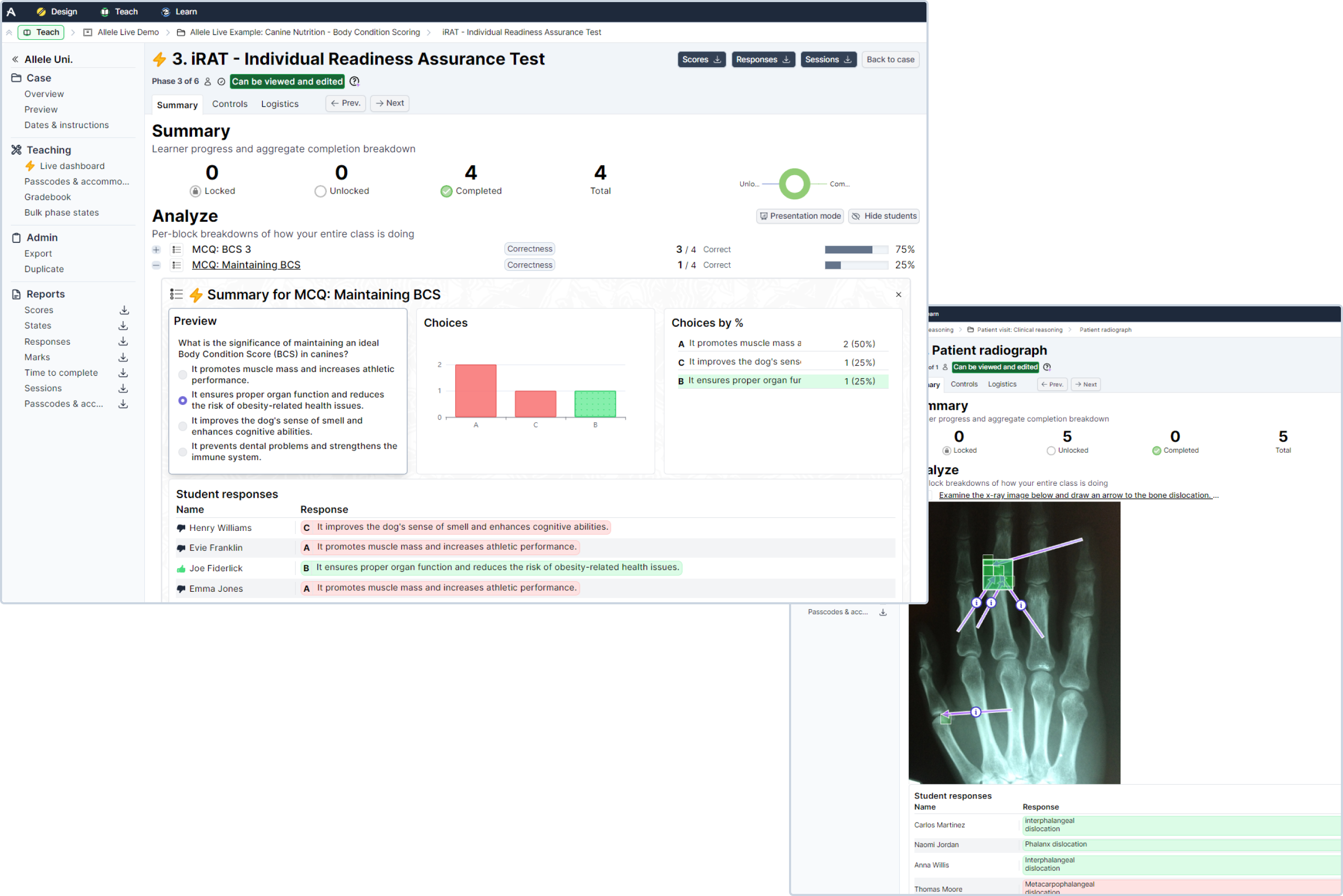
Response Analytics Feature Allele This repository provides a framework for fine grained prediction of chromatin accessibility from dna sequence, using convnext v2 blocks as powerful feature extractors. The report provides summary statistics to evaluate results per sample and per locus (with the possibility to provide a tsv file with loci annotations to include on a table). For this purpose, we developed the maxx (mutation allelic expression extractor) software, which is highly effective at delineating the allelic expression of both single nucleotide variants and small insertions and deletions. In this article, we reviewed methods for detecting ase using both bulk rnaseq and single cell rnaseq data to provide deeper insights beyond the mere prediction of ase genes. fundamentally, ase detection methods are data driven and can be classified according to type of data used.
Comments are closed.