Dna Testing Reveals Hidden Diversity In Ocean Microbiomes

Oceandna Genetic Reseach Section Aori Utokyo Here we recovered 43,191 bacterial and archaeal genomes from publicly available marine metagenomes, encompassing a wide range of diversity with 138 distinct phyla, redefining the upper limit. The breakthrough technique will help researchers to diagnose conditions at the base of the ocean food web that affects the abundance of commercially important fishes or creates harmful algal blooms.
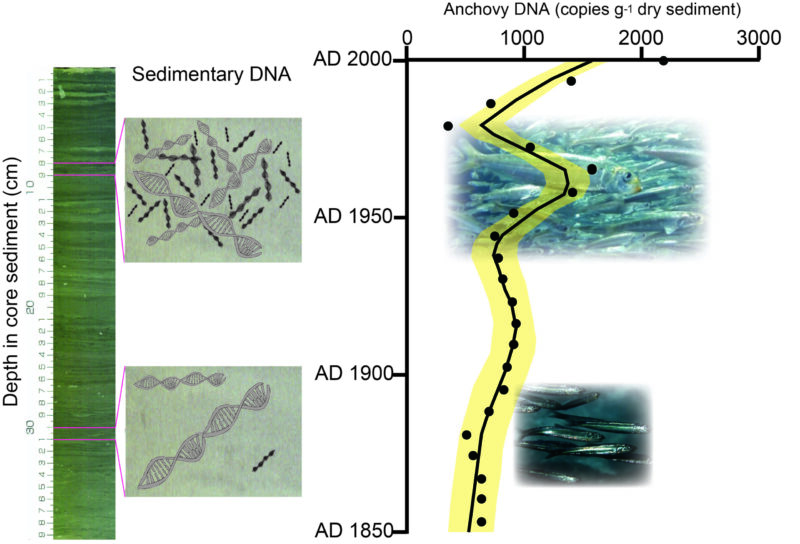
The First Detection Of Marine Fish Dna In Sediment Sequences Going Back In a new study in nature microbiology, researchers from bigelow laboratory for ocean sciences and atrandi biosciences provide the first environmental application of the approach, which they call. Researchers used dna analysis to comprehensively map microbial ocean life off the california coast. learn about the findings from scripps, jcvi & noaa. This forum focuses on the vast diversity of deep sea microbes and their potential for bioprospecting. it also discusses threats posed by climate change and deep sea mining to deep sea microbial genetic resources, and proposes future research directions. This is particularly true for the deep sea meiofauna and eukaryotic microbiota, whose hidden diversity is largely unexplored. here, we tackle this issue by using unique dna signatures to classify unknown metabarcodes assigned to deep sea foraminifera.

Dna Testing Reveals Hidden Diversity In Ocean Microbiomes This forum focuses on the vast diversity of deep sea microbes and their potential for bioprospecting. it also discusses threats posed by climate change and deep sea mining to deep sea microbial genetic resources, and proposes future research directions. This is particularly true for the deep sea meiofauna and eukaryotic microbiota, whose hidden diversity is largely unexplored. here, we tackle this issue by using unique dna signatures to classify unknown metabarcodes assigned to deep sea foraminifera. Scientists at the scripps institution of oceanography and other institutions have used tools of genetics akin to those in genealogical research to evaluate the diversity of marine life off the california coast. Environmental dna (edna) metabarcoding is a promising and cost efficient biodiversity monitoring tool with great potential to uncover the hidden genetic diversity and assess the state and functioning of marine ecosystems. In the case of this study, researchers were able to use genetic information to identify the most important factor governing how many organisms are in the ocean in surface waters off the california coast and where they are distributed. Fungi represented over 50% of the distinct gene clusters identified in the mesopelagic zone. this finding highlights the contribution of fungi to microbial diversity and carries functional consequences for the role of fungi in elemental cycling in the ocean.

Comprehensive Regional Diagnostic Of Microbial Ocean Life Using Dna Scientists at the scripps institution of oceanography and other institutions have used tools of genetics akin to those in genealogical research to evaluate the diversity of marine life off the california coast. Environmental dna (edna) metabarcoding is a promising and cost efficient biodiversity monitoring tool with great potential to uncover the hidden genetic diversity and assess the state and functioning of marine ecosystems. In the case of this study, researchers were able to use genetic information to identify the most important factor governing how many organisms are in the ocean in surface waters off the california coast and where they are distributed. Fungi represented over 50% of the distinct gene clusters identified in the mesopelagic zone. this finding highlights the contribution of fungi to microbial diversity and carries functional consequences for the role of fungi in elemental cycling in the ocean.

Environmental Dna Left Behind By Ocean Animals Can Measure Marine In the case of this study, researchers were able to use genetic information to identify the most important factor governing how many organisms are in the ocean in surface waters off the california coast and where they are distributed. Fungi represented over 50% of the distinct gene clusters identified in the mesopelagic zone. this finding highlights the contribution of fungi to microbial diversity and carries functional consequences for the role of fungi in elemental cycling in the ocean.

Dna Left By Ocean Animals Provides Rare Glimpse Of Marine Ecosystems
Comments are closed.