Bioinformatics Par 14 Dot Plot
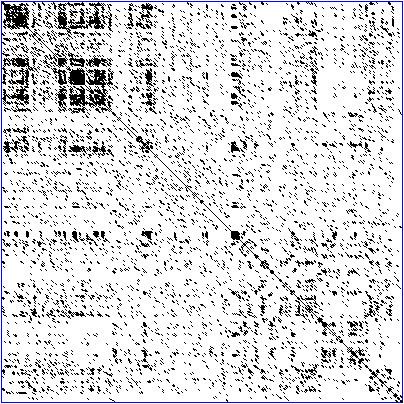

Dot Plot Bioinformatics Alchetron The Free Social Encyclopedia This bioinformatics tutorial explains dot plot and dot matrix analysis of two sequences for the dynamic programming alignment. for more information, log on t. A dot plot is a 2 dimensional matrix where each axis of the plot represents one sequence. by sliding a fixed size window over the sequences and making a sequence match by a dot in the matrix, a diagonal line will emerge if two identical (or very homologous) sequences are plotted against each other.
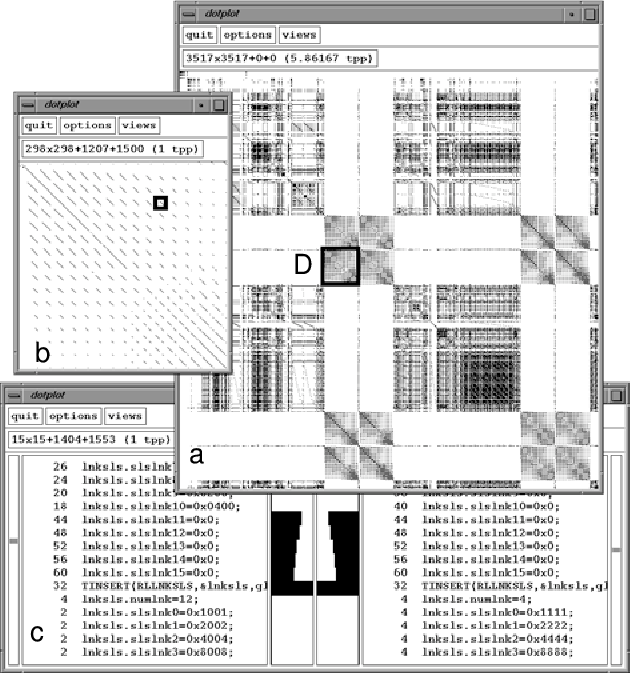

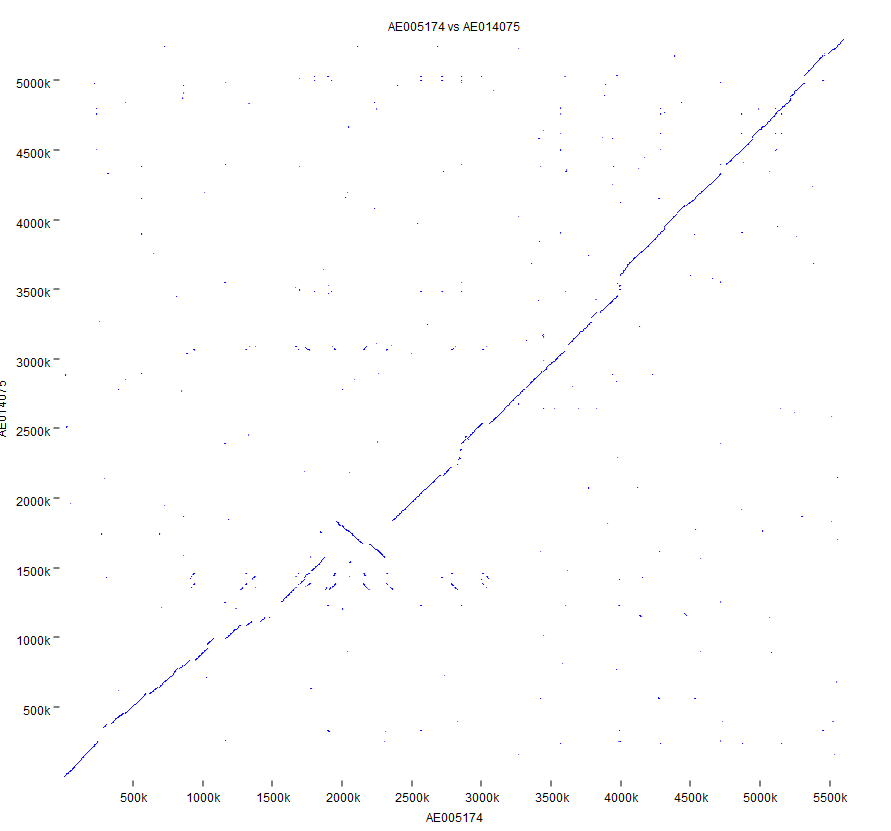
Dot Plot Bioinformatics Semantic Scholar In this hands on session we will explore the principles underlying the computational tools that can be used to compute and evaluate sequence alignments. dot plots are a simple graphical approach for the visual comparison of two sequences. The document discusses how to create dot plots between dna and protein sequences and explains how using a sliding window threshold can filter out random matches. We are going to make dotplots, which is a way to line up two genomes and see high level structural differences between them (large insertions and deletions, translocations, inversions, etc). single nucleotide changes (snps) and small indels are ignored at this level. we’ll start with sars cov 2. For each word, determine a neighborhood of words that, if found in another sequence, would likely to be part of a significant maximum scoring pair (msp). scan databases for neighborhood words.
Bioinformatics Dot Plot Matrix Python Ipynb At Main Prasanth S N We are going to make dotplots, which is a way to line up two genomes and see high level structural differences between them (large insertions and deletions, translocations, inversions, etc). single nucleotide changes (snps) and small indels are ignored at this level. we’ll start with sars cov 2. For each word, determine a neighborhood of words that, if found in another sequence, would likely to be part of a significant maximum scoring pair (msp). scan databases for neighborhood words. Figure 18.11: the dot plot showing a low complexity region in the sequence. the sequence is artificial and low complexity regions do not always show as a square. A dot plot (aka contact plot or residue contact map) is a graphical method that allows the comparison of two biological sequences and identify regions of close similarity between them. A dot plot is a graphical representation used in bioinformatics to visualize the similarity between two biological sequences, typically dna, rna, or protein sequences. Dot plot matrices produced by the dot plot algorithm can be visualized using this function for a variety of tasks including sequence alignment, genome comparison, and motif discovery.
Qiagen Bioinformatics Manuals Figure 18.11: the dot plot showing a low complexity region in the sequence. the sequence is artificial and low complexity regions do not always show as a square. A dot plot (aka contact plot or residue contact map) is a graphical method that allows the comparison of two biological sequences and identify regions of close similarity between them. A dot plot is a graphical representation used in bioinformatics to visualize the similarity between two biological sequences, typically dna, rna, or protein sequences. Dot plot matrices produced by the dot plot algorithm can be visualized using this function for a variety of tasks including sequence alignment, genome comparison, and motif discovery.

Introduction To Bioinformatics Dot Plots Dot Plots One A dot plot is a graphical representation used in bioinformatics to visualize the similarity between two biological sequences, typically dna, rna, or protein sequences. Dot plot matrices produced by the dot plot algorithm can be visualized using this function for a variety of tasks including sequence alignment, genome comparison, and motif discovery.
Github Longpham7 Biological Sequences Dot Plot Dot Matrix Analysis
Comments are closed.