Anz Pacbio Workshop Series Day 2 Genome Annotation
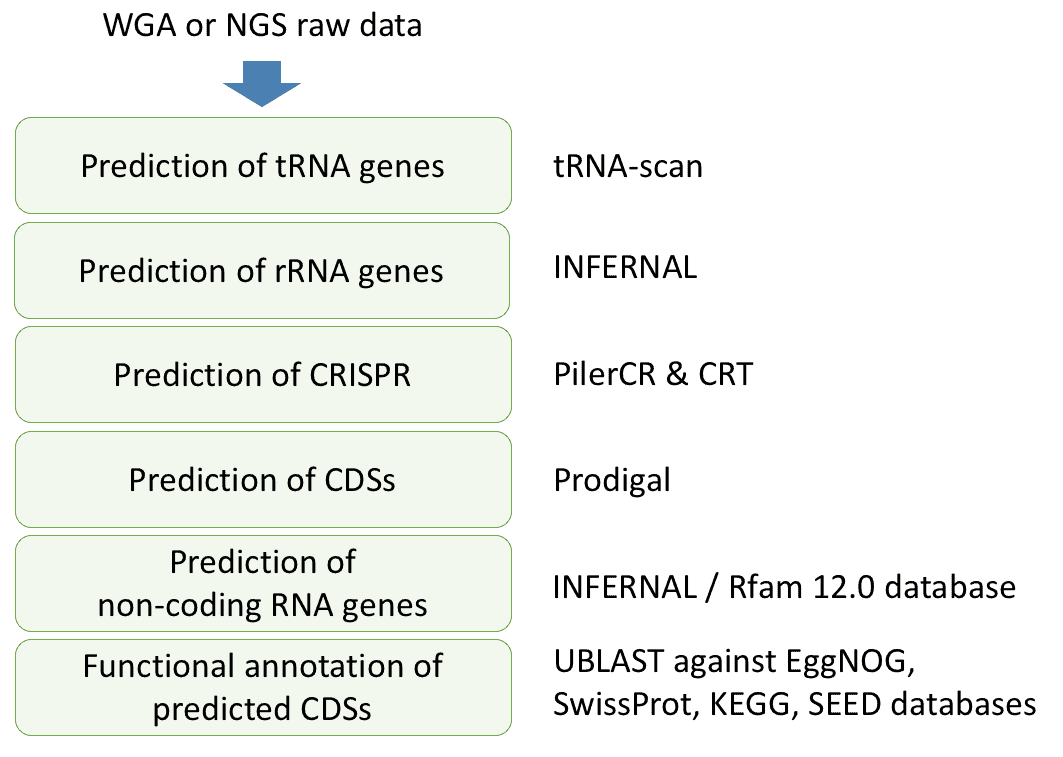

Genome Annotation Ezbiocloud Help Center Catch up with the recording of day #2 of our pacbio workshop series, with this webinar on genome annotation. thanks to pacbio and millennium science for this fun collaboration!. This technical training presentation describes workflow details for constructing whole genome sequencing (wgs) libraries from genomic and metagenomic dna using smrtbell prep kit 3.0 (spk 3.0) for sequencing on pacbio sequel ii, sequel iie and revio systems.

Github Galikbence Genome Annotation Pipeline For Eukaryotic Genome Contribute to ebp nor workshop 2024 development by creating an account on github. Over the past two decades, advances in high throughput sequencing techniques have enabled faster and more accurate capture of diverse genomic features, generating data at an unprecedented scale. And we're off, with day #2 of our anz pacbio bioinformatics workshop! up first we have dr paul gooding of millennium science, opening with his talk. This database can consist of one single well annotated reference genome, the whole phage catalog of ncbi or a set of well curated protein annotations such as phrogs.
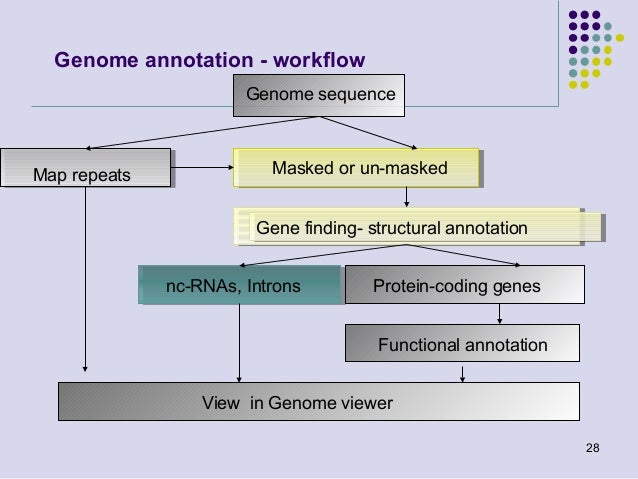
Genome Annotation 2013 And we're off, with day #2 of our anz pacbio bioinformatics workshop! up first we have dr paul gooding of millennium science, opening with his talk. This database can consist of one single well annotated reference genome, the whole phage catalog of ncbi or a set of well curated protein annotations such as phrogs. Unlock the secrets of genome annotation with this comprehensive, hands on virtual workshop! whether you're new to bioinformatics or want to refine your skills, this workshop will guide you through linux, genome assembly, annotation with various types of evidence, and evaluation. This two day in person workshop provides a structured approach to assembling genomes from long read sequencing data (pacbio, ont) and annotating gene models using state of the art computational tools. Some of the best practices and expectations to consider for microbial genome sequencing with pacbio are: start with the recommended input of highquality dna (1.0 μg) per genome. Workshop participants will gain knowledge and hands on experience in analysing pacbio hifi sequencing data, running whole genome assemblies, evaluating the quality of their genome assemblies, and will be introduced to whole genome annotation.

Upcoming Virtual Genome Annotation Workshop Register Now Unlock the secrets of genome annotation with this comprehensive, hands on virtual workshop! whether you're new to bioinformatics or want to refine your skills, this workshop will guide you through linux, genome assembly, annotation with various types of evidence, and evaluation. This two day in person workshop provides a structured approach to assembling genomes from long read sequencing data (pacbio, ont) and annotating gene models using state of the art computational tools. Some of the best practices and expectations to consider for microbial genome sequencing with pacbio are: start with the recommended input of highquality dna (1.0 μg) per genome. Workshop participants will gain knowledge and hands on experience in analysing pacbio hifi sequencing data, running whole genome assemblies, evaluating the quality of their genome assemblies, and will be introduced to whole genome annotation.
Comments are closed.