3d Genomics For Structural Variant Detection Technology Update

Application Note Structural Variant Detection With Hi C Technology New experimental and computational approaches are revealing the effect of structural variants on the 3d genome and gene expression and can help interpret their pathogenic potential, which. Each technology has provided genome wide access to different classes of human genetic variation. we are now on the verge of comprehensive variant detection of all forms of variation for the first time with a single assay.
Github Roboatory Structural Variant Detection Structural Variant We utilized a multi purpose ngs assay (dovetail® linkprep™) to detect snvs, indels, cnvs, svs and 3d genome contact maps. peripheral blood or bone marrow cells were thawed and immediately fixed, subjected to crosslinking and in situ tagmentation, followed by library prep. Clinical cancer sequencing also increasingly aims to detect svs, leading to a widespread necessity to interpret them. recently, analyses of large whole genome sequencing datasets revealed features that impact rates of svs across the genome in different cancers. To investigate the effects of variants in the context of 3d organization, a two phase scoring algorithm, 3dfunc, is developed to evaluate the pathogenicity of variant–gene pairs in cancer. By integrating sequence data with spatial genome mapping, researchers can detect structural variants, enhancer hijacking events, and other regulatory disruptions while interpreting their biological consequences.
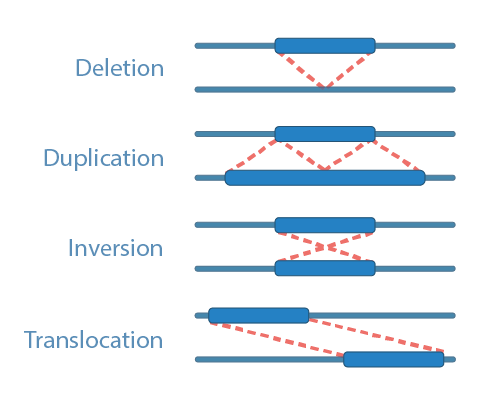
Structural Variant Analysis To investigate the effects of variants in the context of 3d organization, a two phase scoring algorithm, 3dfunc, is developed to evaluate the pathogenicity of variant–gene pairs in cancer. By integrating sequence data with spatial genome mapping, researchers can detect structural variants, enhancer hijacking events, and other regulatory disruptions while interpreting their biological consequences. While traditional short read sequencing is limited in detecting structural variants, combining high throughput chromatin conformation capture (hi c) technology with short read sequence data enables the detection of svs5. In this work, we introduce structural variants genotyping of assemblies on population scales (svgap), a flexible pipeline to detect, genotype, and annotate svs in large samples of de novo genome assemblies. Standard laboratory tests can fail to detect many disease causing dna changes. now, a novel 3d chromosome mapping method can reliably reveal these hidden structural variants and lead to new. New experimental and computational approaches are revealing the effect of structural variants on the 3d genome and gene expression and can help interpret their pathogenic potential, which has important implications for patients.

Advanced Structural Variant Detection Free Plugin Bioinformatics While traditional short read sequencing is limited in detecting structural variants, combining high throughput chromatin conformation capture (hi c) technology with short read sequence data enables the detection of svs5. In this work, we introduce structural variants genotyping of assemblies on population scales (svgap), a flexible pipeline to detect, genotype, and annotate svs in large samples of de novo genome assemblies. Standard laboratory tests can fail to detect many disease causing dna changes. now, a novel 3d chromosome mapping method can reliably reveal these hidden structural variants and lead to new. New experimental and computational approaches are revealing the effect of structural variants on the 3d genome and gene expression and can help interpret their pathogenic potential, which has important implications for patients.
Comments are closed.