Visualizing Gene Expression
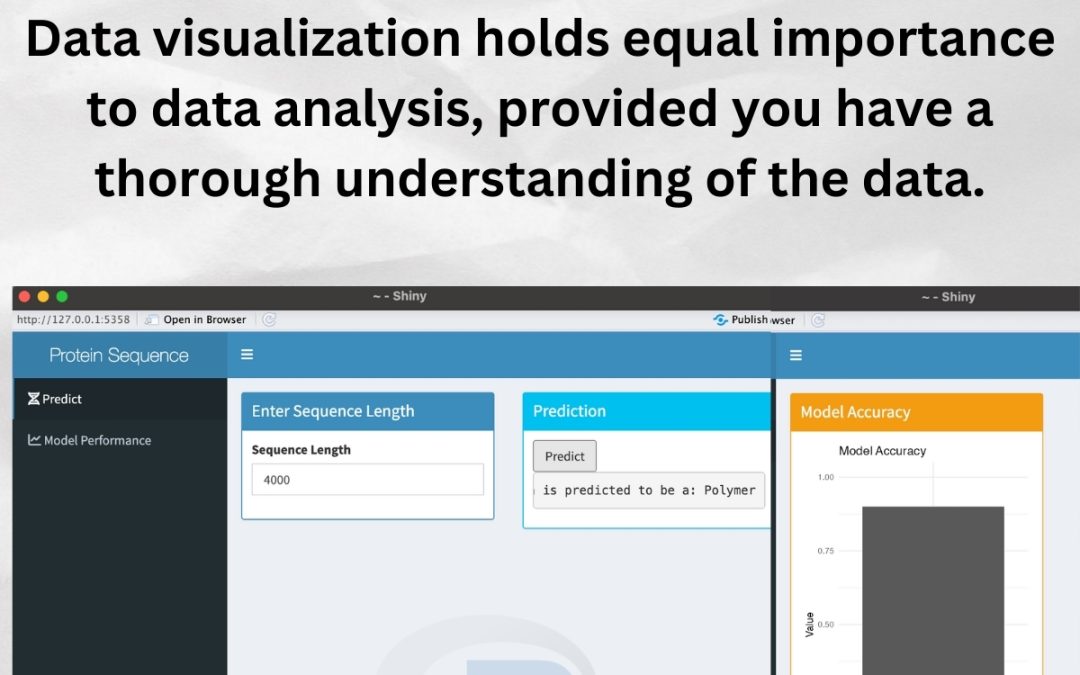
A Dynamic Dashboard Visualizing Gene Expression Data Aiinbioinformatics To improve the flexibility in handling different data structures, expressyourcell provides the user with multiple options for coloring subcellular localizations (see details in section visualizing gene expression data to the cellular pictograph for details). To improve the flexibility in handling different data structures, expressyourcell provides the user with multiple options for coloring subcellular localizations (see details in section visualizing gene expression data to the cellular pictograph for details).
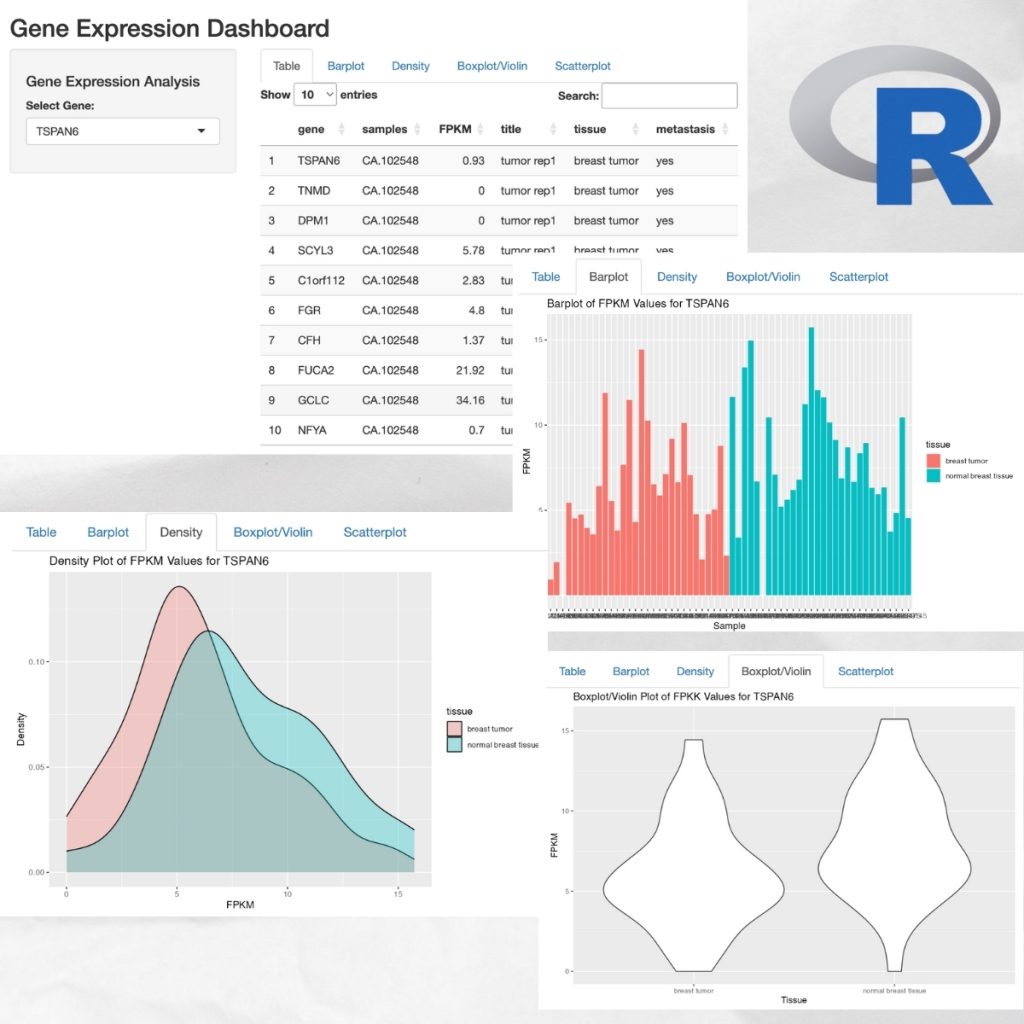
A Dynamic Dashboard Visualizing Gene Expression Data Aiinbioinformatics Visualizing gene expression patterns | this tutorial describes methods used to detect when and where genes are expressed in cells and tissues. In this tutorial, you will learn how to: compute and visualize gene expression trends along specific differentiation trajectories. cluster expression trends to detect genes with similar dynamical patterns. this tutorial notebook can be downloaded using the following link. In this paper, we 1) introduce new interactive rna seq visualization tools, 2) compile a collection of examples that demonstrate to biologists why visualization should be an integral component of differential expression analysis. Phantasus democratizes gene expression analysis, offering intuitive and interactive tools that streamline the exploration and analysis of user provided data and over 96,000 public datasets.
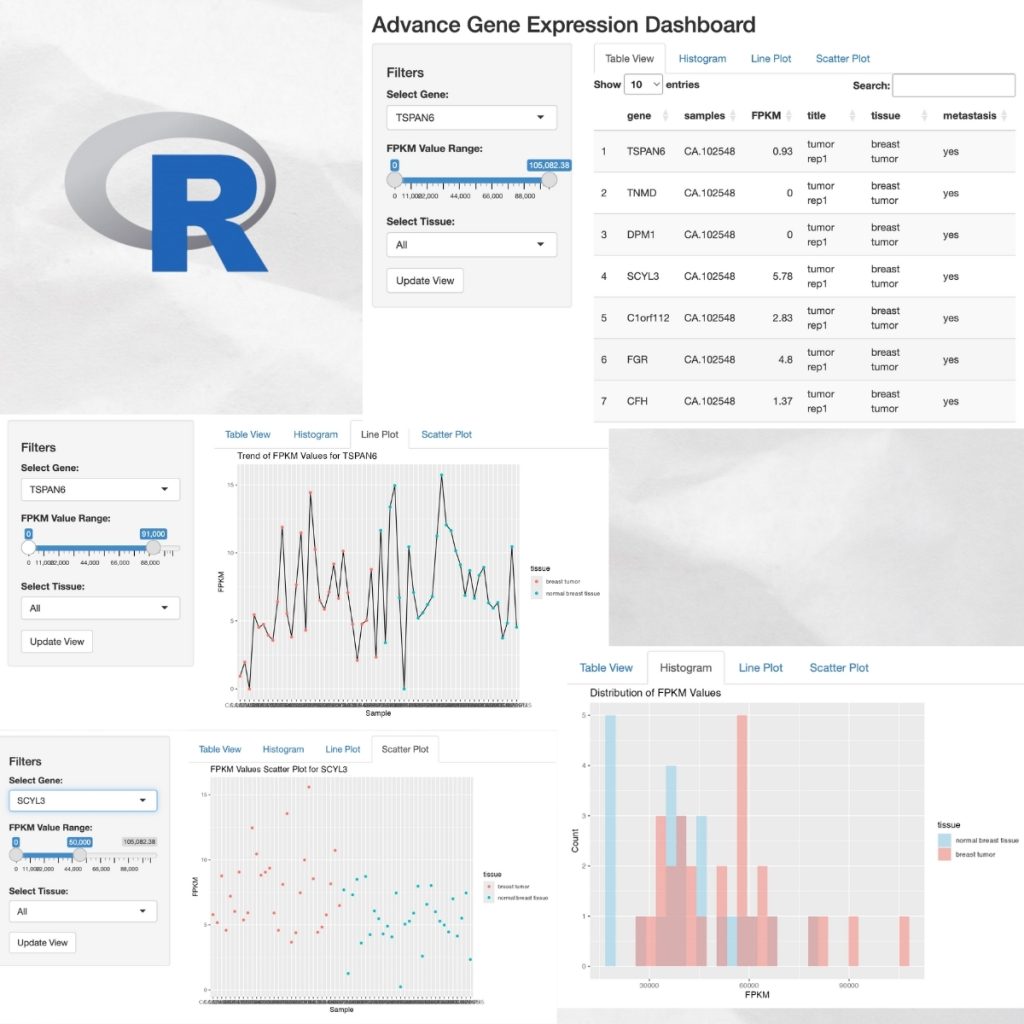
A Dynamic Dashboard Visualizing Gene Expression Data Aiinbioinformatics In this paper, we 1) introduce new interactive rna seq visualization tools, 2) compile a collection of examples that demonstrate to biologists why visualization should be an integral component of differential expression analysis. Phantasus democratizes gene expression analysis, offering intuitive and interactive tools that streamline the exploration and analysis of user provided data and over 96,000 public datasets. Execute interactive analyses on a dataset to explore how patterns of gene expression are determined by spatial, environmental, and genetic factors using an interactive speed no code ui. With this visualization we can appreciate the spatial localization of gene expression changes and observe the emergence of downregulated genes at 30 min, with robust negative fold changes in mrnas encoding for proteins localized in the endoplasmic reticulum. Heatmaps are excellent for visualizing expression patterns across multiple genes and samples. they can reveal gene clusters and expression patterns that might not be obvious from other visualizations. Embl ebi expression atlas, an open public repository of gene expression pattern data under different biological conditions.

Visualizing Gene Expression With Mri Www Caltech Edu Execute interactive analyses on a dataset to explore how patterns of gene expression are determined by spatial, environmental, and genetic factors using an interactive speed no code ui. With this visualization we can appreciate the spatial localization of gene expression changes and observe the emergence of downregulated genes at 30 min, with robust negative fold changes in mrnas encoding for proteins localized in the endoplasmic reticulum. Heatmaps are excellent for visualizing expression patterns across multiple genes and samples. they can reveal gene clusters and expression patterns that might not be obvious from other visualizations. Embl ebi expression atlas, an open public repository of gene expression pattern data under different biological conditions.

Visualizing Gene Expression With Mri Lab Manager Heatmaps are excellent for visualizing expression patterns across multiple genes and samples. they can reveal gene clusters and expression patterns that might not be obvious from other visualizations. Embl ebi expression atlas, an open public repository of gene expression pattern data under different biological conditions.
Comments are closed.