Varseq Variantanalysisresearch

Utah State University Bioinformatics Facility From raw vcf to clinically actionable findings, varseq delivers an integrated variant analysis workflow with real time filter chains, curated annotation sources, and powerful prioritization across panels, exomes, and genomes. This document discusses the use of varseq to enhance variant analysis research, detailing its capabilities for variant annotation, filtering, and ranking through a flexible and scalable interface.
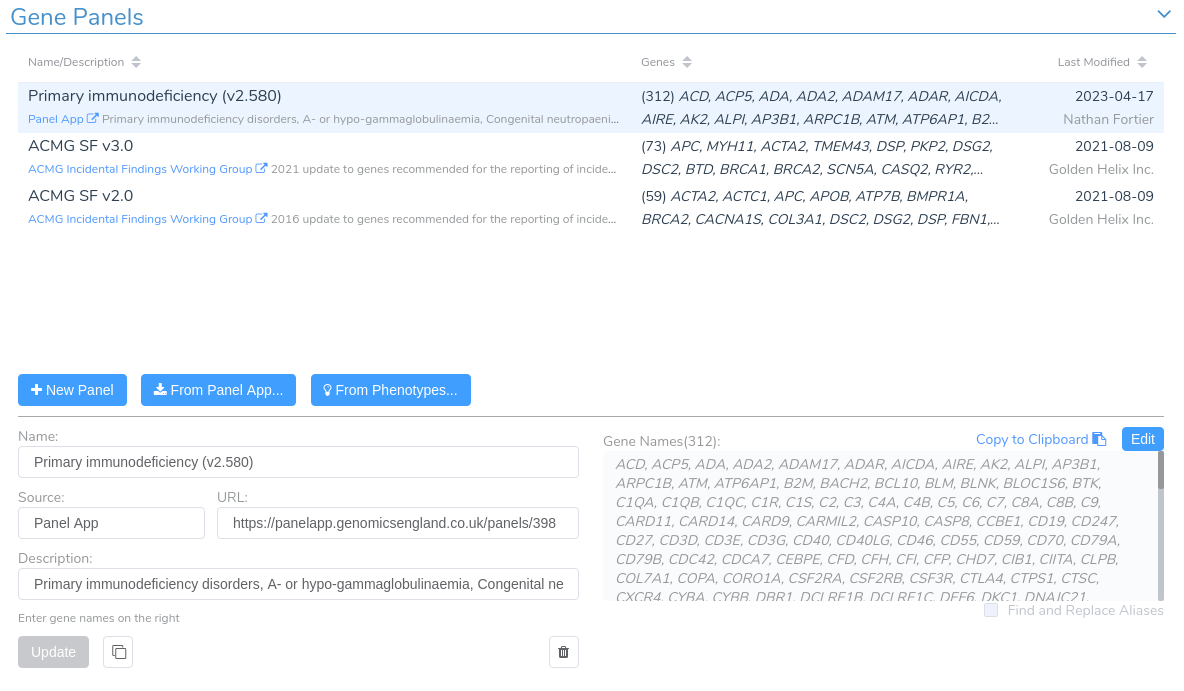
Unraveling Structural Variants With Varseq In this webinar presentation, we will review workflow strategies for quality control and analysis of dna sequence variants using the varseq software package from golden helix. Golden helix has released an updated version of its variant analysis software, varseq 2.6.2. the new version has enhanced pharmacogenomics analysis capabilities, including a cyp2d6 star allele caller called cypcall. Varseq is an innovative software platform that streamlines the analysis of genomic data, ranging from gene panels to whole exomes and genomes. it features an intuitive interface that allows users to automate workflows, enhancing the efficiency and accessibility of variant analysis. Varseq software provides a powerful filtering and annotation engine to sift through large variant data sets. using a chain of filters, you can quickly narrow your list of variants down to those that are most likely to be of interest.
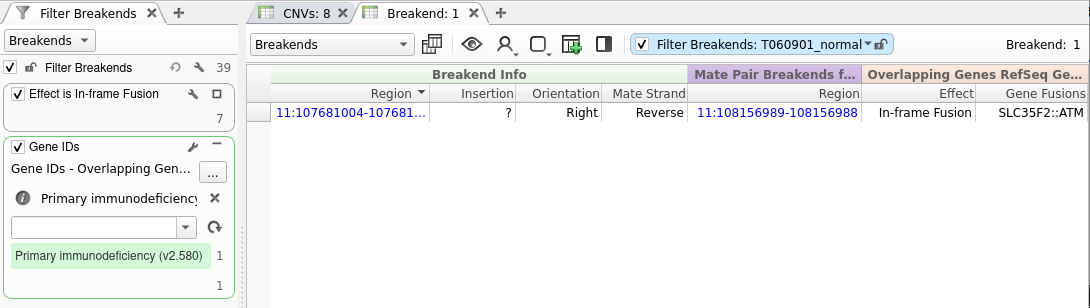
Unraveling Structural Variants With Varseq Varseq is an innovative software platform that streamlines the analysis of genomic data, ranging from gene panels to whole exomes and genomes. it features an intuitive interface that allows users to automate workflows, enhancing the efficiency and accessibility of variant analysis. Varseq software provides a powerful filtering and annotation engine to sift through large variant data sets. using a chain of filters, you can quickly narrow your list of variants down to those that are most likely to be of interest. With varseq, researchers can tackle the intricacies of genomic data more easily, facilitating a straightforward navigation and understanding of their findings. the application includes a powerful filtering and annotation system that allows for the efficient management of large variant datasets. With varseq, the complexities of genomic data become more manageable, enabling researchers to easily navigate and interpret results. the software features a robust filtering and annotation system that helps users efficiently process extensive variant datasets. This webcast demonstrated how varseq integrates genomenon mastermind and genomenon cancer knowledgebase (ckb), enhancing germline and somatic variant analysis and clinical reporting. Varseq is an intuitive, integrated software solution for tertiary analysis. with varseq you can automate your workflows and analyze variants for gene panels, exomes, and whole genomes.
Comments are closed.