Variant Curation Documents Update
Rare Source Home A list of resources related to the process of curating, assessing and classifying a variant through a clingen approved variant curation expert panel (vcep). updates to these resources are versioned over time and the most updated version is available on the website. In this video, the variant curation proxies provides updates on the vcep protocol and vcep evidence summary standardized text documents, as well as the new points feature on the criteria.
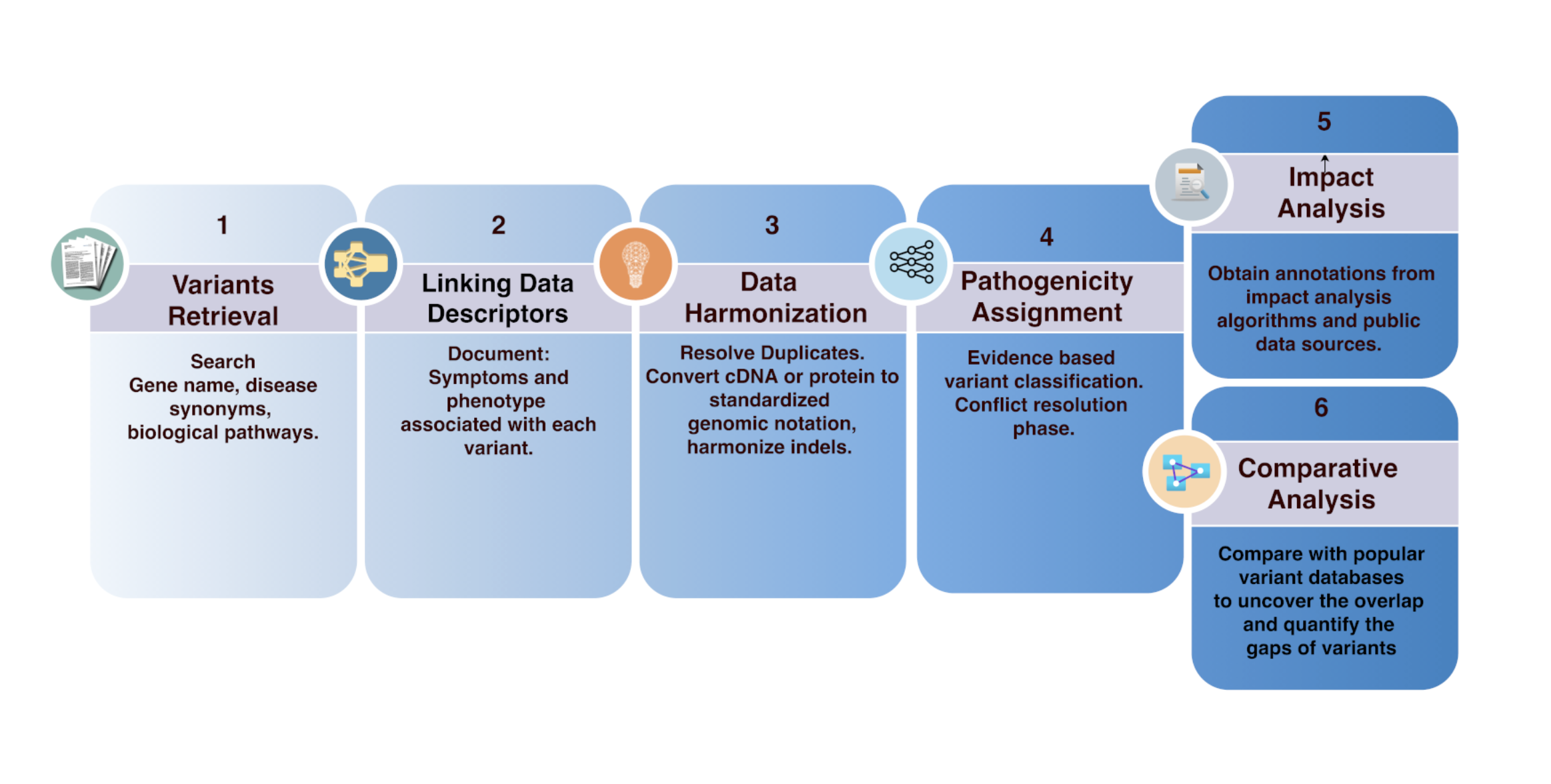
Clingen Variant Curation Expert Panel Experiences And Standardized This updated version of the acgs guidelines also includes guidelines for the reporting of non coding sequence variants, reduced penetrance, hypomorphic sequence variants and risk alleles. it is essential that clinical scientists use their professional judgement for variant classification. To facilitate clinvar submission with the increasing growth in clingen curations, we have developed the data model and user workflow in the vci to support affiliations submitting variant curations and classifications to clinvar using an api. Here you can find resources that are either critical for curation or useful for interpretation. there are also descriptions of how to do a thorough literature search and how we organize our. Ldh provides permalinks to timestamped excerpts to document supporting evidence at the time of variant curation or classification. it supports versioning and update propagation that retains information about earlier records from the same source.

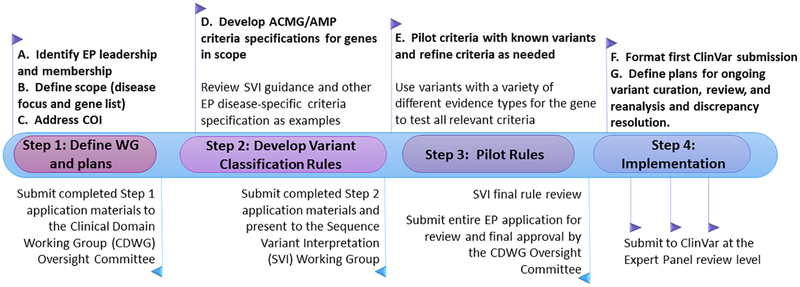
Docm A Database Of Curated Mutations In Cancer Nature Methods Here you can find resources that are either critical for curation or useful for interpretation. there are also descriptions of how to do a thorough literature search and how we organize our. Ldh provides permalinks to timestamped excerpts to document supporting evidence at the time of variant curation or classification. it supports versioning and update propagation that retains information about earlier records from the same source. Updates to files for download xml files. the clinvar xml files were updated to allow distinct classifications in the germline and somatic contexts. in the old xml format there was a single element, clinicalsignificance, for the classification of the variant. The original population guidelines were used as a case study to assess the necessary parameters and level of evidence for each threshold. we thoroughly examined the main disease causing variants found by each vcep in their guidance document while evaluating their initial variant classification to confirm the thresholds they established. These gene specific rules for atm were modified from the american college of medical genetics and association for molecular pathology (acmg amp) guidelines and were tested against 33 atm variants of various types and classifications in a pilot curation phase. Cdh1 variant classification specifications were modified based on updated genetic testing clinical criteria, new recommendations from clingen, and expert knowledge from ongoing cdh1 variant curations.

Variant Level Matching For Diagnosis And Discovery Challenges And Updates to files for download xml files. the clinvar xml files were updated to allow distinct classifications in the germline and somatic contexts. in the old xml format there was a single element, clinicalsignificance, for the classification of the variant. The original population guidelines were used as a case study to assess the necessary parameters and level of evidence for each threshold. we thoroughly examined the main disease causing variants found by each vcep in their guidance document while evaluating their initial variant classification to confirm the thresholds they established. These gene specific rules for atm were modified from the american college of medical genetics and association for molecular pathology (acmg amp) guidelines and were tested against 33 atm variants of various types and classifications in a pilot curation phase. Cdh1 variant classification specifications were modified based on updated genetic testing clinical criteria, new recommendations from clingen, and expert knowledge from ongoing cdh1 variant curations.
Comments are closed.