Variant Calling Using Bcftools Omicsbox User Manual
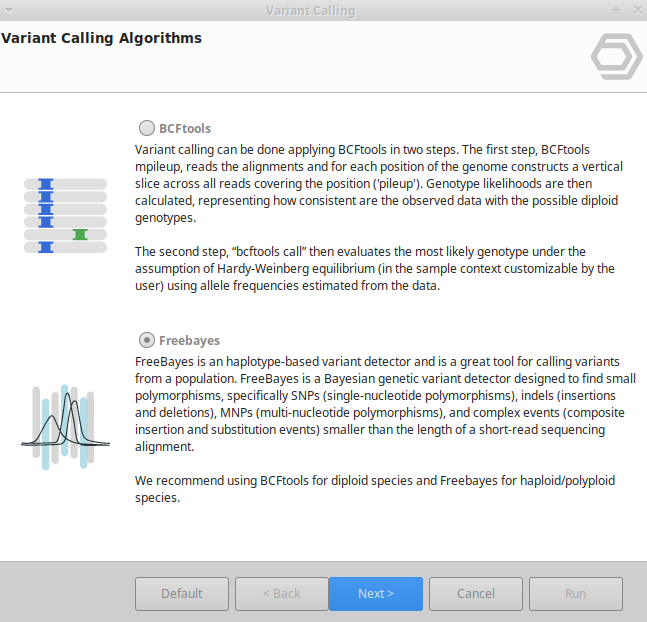
Variant Calling Omicsbox User Manual Identify genetic variants with bcftools in omicsbox, using advanced algorithms for mpileup and genotype calling with customizable quality parameters. Bcftools is a widely used variant calling tool, especially among non human species, which is characterized by its small time of execution and its precision. bcftools uses two algorithms.
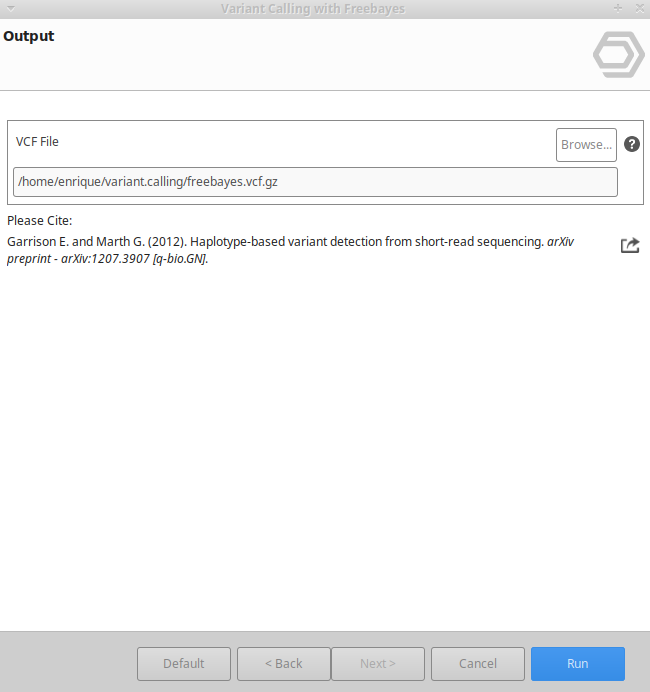
Variant Calling Using Freebayes Omicsbox User Manual Bcftools functions effectively as a sibling package to samtools, heavily targeted toward manipulating and generating vcf (variant call format) representations. within modern workflows, it is universally used to call single nucleotide variants (snvs), define critical indels, and output detailed consensus strings. One can get a feel for typical distributions of annotations from a set of known good calls vs all calls. if you don’t already have a pipeline for this, use bcftools query to extract the annotations and plot manually, for example like this:. First let's see how to use a simple pipeline to identify genetic variants using bcftools mpileup and bcftools call. as this suggests the process has two steps. in the first step (the mpileup step), we process the reads, identify likely alleles, and compute genotype likelihoods. Bcftools is a set of utilities that manipulate variant calls in the variant call format (vcf) and its binary counterpart bcf. all commands work transparently with both vcfs and bcfs, both uncompressed and bgzf compressed.
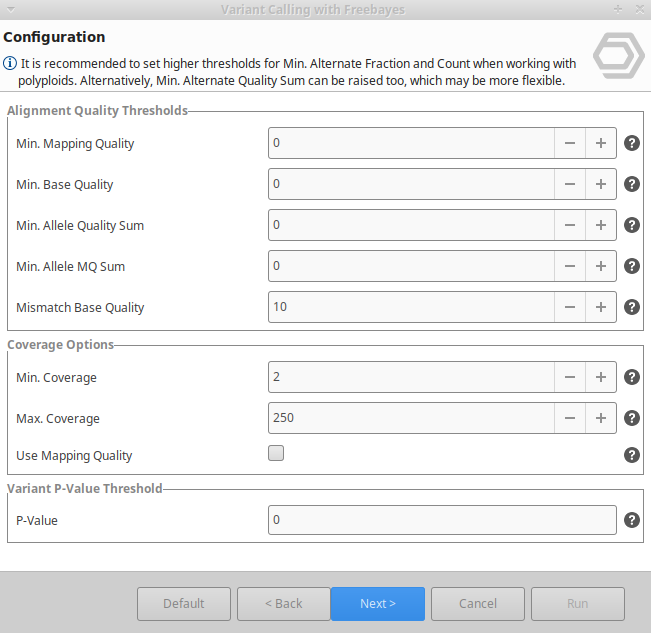
Variant Calling Using Freebayes Omicsbox User Manual First let's see how to use a simple pipeline to identify genetic variants using bcftools mpileup and bcftools call. as this suggests the process has two steps. in the first step (the mpileup step), we process the reads, identify likely alleles, and compute genotype likelihoods. Bcftools is a set of utilities that manipulate variant calls in the variant call format (vcf) and its binary counterpart bcf. all commands work transparently with both vcfs and bcfs, both uncompressed and bgzf compressed. In this tutorial we will first do variant calling using bcftools, and then we will explore the resulting files a bit. Bcftools is a set of utilities that manipulate variant calls in the variant call format (vcf) and its binary counterpart bcf. all commands work transparently with both vcfs and bcfs, both uncompressed and bgzf compressed. Similar to other steps in this workflow, there are a number of tools available for variant calling. in this workshop we will be using bcftools, but there are a few things we need to do before actually calling the variants. This page describes the variant calling pipeline in bcftools, which identifies snps and indels from aligned sequence reads (bam cram files).
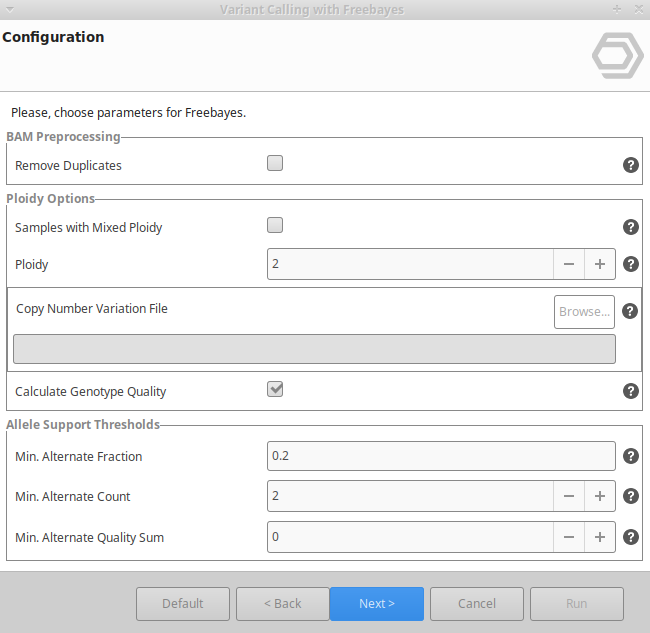
Variant Calling Using Freebayes Omicsbox User Manual In this tutorial we will first do variant calling using bcftools, and then we will explore the resulting files a bit. Bcftools is a set of utilities that manipulate variant calls in the variant call format (vcf) and its binary counterpart bcf. all commands work transparently with both vcfs and bcfs, both uncompressed and bgzf compressed. Similar to other steps in this workflow, there are a number of tools available for variant calling. in this workshop we will be using bcftools, but there are a few things we need to do before actually calling the variants. This page describes the variant calling pipeline in bcftools, which identifies snps and indels from aligned sequence reads (bam cram files).
Variant Calling Using Freebayes Omicsbox User Manual Similar to other steps in this workflow, there are a number of tools available for variant calling. in this workshop we will be using bcftools, but there are a few things we need to do before actually calling the variants. This page describes the variant calling pipeline in bcftools, which identifies snps and indels from aligned sequence reads (bam cram files).
Variant Calling Using Freebayes Omicsbox User Manual
Comments are closed.