Transcriptomics Tutorial New

Spatial Transcriptomics New Year New Technology Bioinformatics Enjoy the videos and music you love, upload original content, and share it all with friends, family, and the world on . In addition to infer the intercellular communication from any given scrna seq data and spatially resolved transcriptomics data, cellchat provides functionality for further data exploration, analysis, and visualization. it can quantitatively characterize and compare the inferred cell cell communication networks using an integrated approach by combining social network analysis, pattern.

Wikipathways Publications Transcriptomics Analysis Reveals New Training material for all kinds of transcriptomics analysis. before diving into this topic, we recommend you to have a look at: you can view the tutorial materials in different languages by clicking the dropdown icon next to the slides () and tutorial () buttons below. start here if you are new to rna seq analysis in galaxy. Teaching students how to use open source tools to analyze rnaseq data since 2015. Explain the difference between population genomics and transcriptomics. evaluate how mapping and the “reference” differs for population genomics and transcriptomics. Learn all about illumina's 'sequencing by synthesis' technology, and the steps involved in planning for a transcriptomics experiment. in the first half of this lecture we'll discuss the open source, cross platform r bioconductor software that we will use throughout the course.

New Standards Boost Reliability In Spatial Transcriptomics Technology Explain the difference between population genomics and transcriptomics. evaluate how mapping and the “reference” differs for population genomics and transcriptomics. Learn all about illumina's 'sequencing by synthesis' technology, and the steps involved in planning for a transcriptomics experiment. in the first half of this lecture we'll discuss the open source, cross platform r bioconductor software that we will use throughout the course. Researchers and practitioners must stay abreast with emerging trends, novel technologies, and new analytical methods to leverage the full potential of transcriptomics in biological research. Gain insight into a topic and learn the fundamentals. this course will cover bioinformatics methods for analyzing transcriptomic rna sequencing data generated with the short read (rna seq) and long read (pacbio, ont) sequencing. Tutorial: transcriptome annotation with genemunge, archs4 and mygene.info this notebook demonstrates how to annotate rnalib transcriptome features with annotations from public databases using. We outline a computational workflow that includes quality control, trimming of reads, read alignment to the genome, and gene quantification, ultimately enabling researchers to identify differentially expressed genes and gain insights on mrna levels.
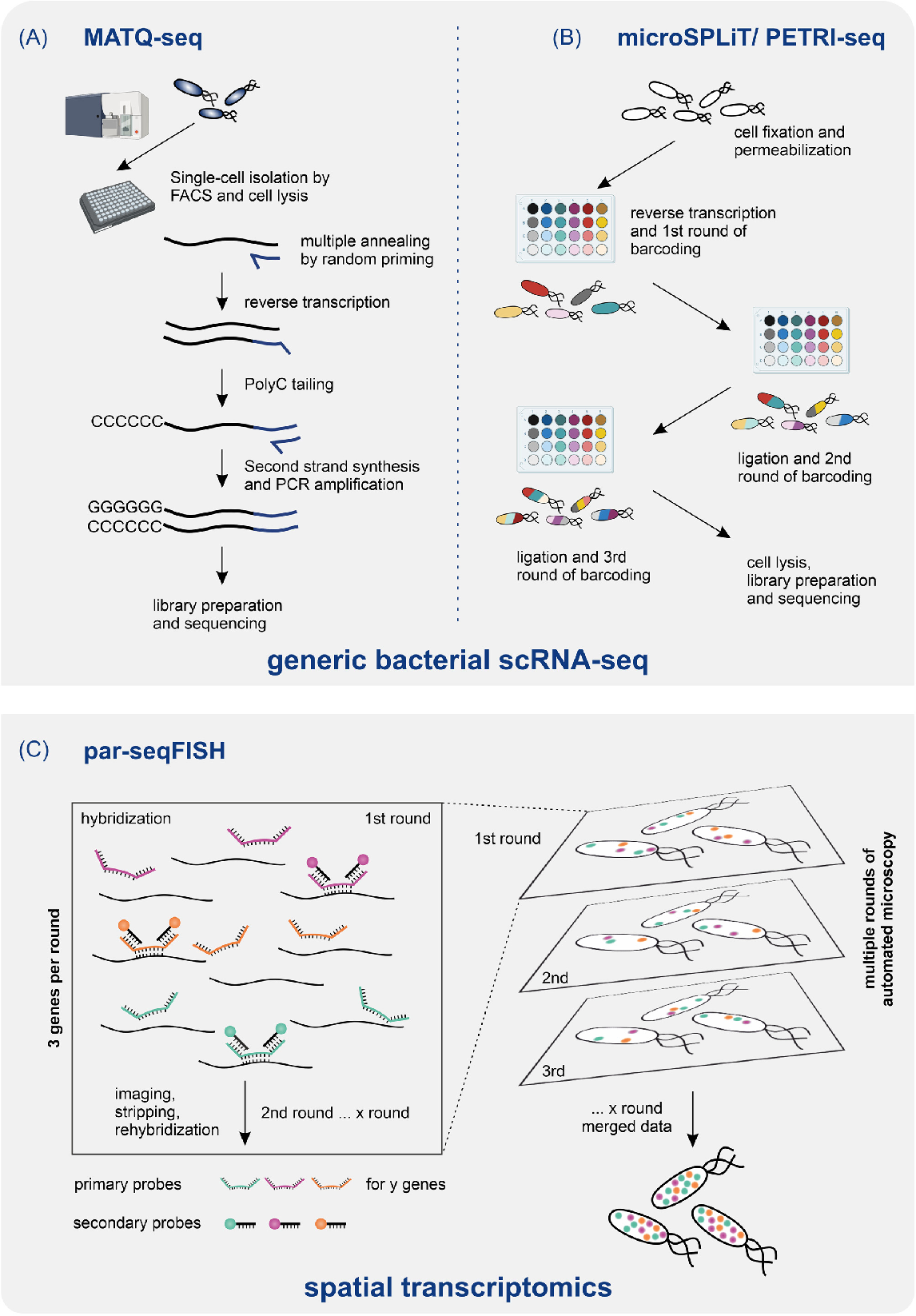
Figure 2 From Ushering In A New Era Of Single Cell Transcriptomics In Researchers and practitioners must stay abreast with emerging trends, novel technologies, and new analytical methods to leverage the full potential of transcriptomics in biological research. Gain insight into a topic and learn the fundamentals. this course will cover bioinformatics methods for analyzing transcriptomic rna sequencing data generated with the short read (rna seq) and long read (pacbio, ont) sequencing. Tutorial: transcriptome annotation with genemunge, archs4 and mygene.info this notebook demonstrates how to annotate rnalib transcriptome features with annotations from public databases using. We outline a computational workflow that includes quality control, trimming of reads, read alignment to the genome, and gene quantification, ultimately enabling researchers to identify differentially expressed genes and gain insights on mrna levels.

Introduction To Transcriptomics Youtube Tutorial: transcriptome annotation with genemunge, archs4 and mygene.info this notebook demonstrates how to annotate rnalib transcriptome features with annotations from public databases using. We outline a computational workflow that includes quality control, trimming of reads, read alignment to the genome, and gene quantification, ultimately enabling researchers to identify differentially expressed genes and gain insights on mrna levels.
Comments are closed.