Transcriptomics Data Analysis With Python Machine Learning Tutorial For Researchers

Python Machine Learning Tutorial Data Science Amazing Elearning Pyscenic is a lightning fast python implementation of the scenic pipeline (single cell regulatory network inference and clustering) which enables biologists to infer transcription factors, gene regulatory networks and cell types from single cell rna seq data. Participants will learn about machine learning techniques for unsupervised and supervised analysis of rna seq data, including clustering and classification methods.

Machine Learning For Transcriptomic Data Workshops Rna Seq Blog The single cell rna seq analysis using python course, which focused on the analysis of single cell rna sequencing (scrna seq) data using python and command line tools, ran in february 2025. here, we have made the course materials available for you to access any time. Purpose: this tutorial guides researchers through preprocessing single cell rna seq (scrna seq) data using scanpy, a fast and scalable python toolkit. it covers quality control, normalization, clustering and cell type annotation. While this enriches our knowledge of the potentials and limitations of machine learning in transcriptomics, it also underscores the challenges of applying ml to transcriptomic data, characterised by its remarkable diversity and complexity. Here, we introduce rnalib, a python library designed to facilitate the development of novel bioinformatics analysis methods for interpreting transcriptomics data.
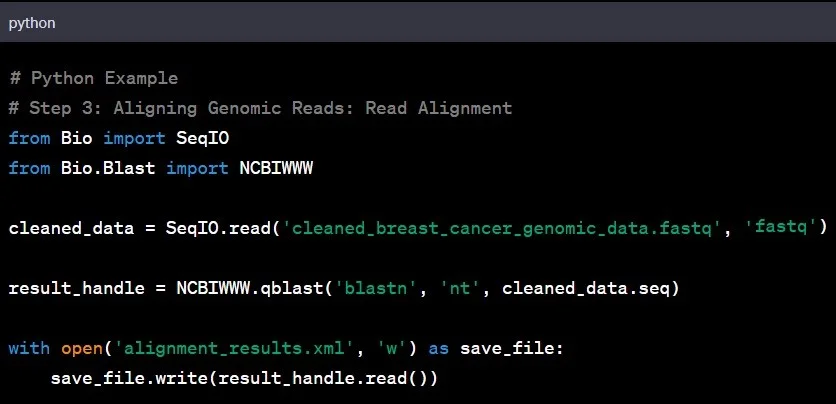
Genomic Data Analysis A Beginner S Step By Step Guide With Python And While this enriches our knowledge of the potentials and limitations of machine learning in transcriptomics, it also underscores the challenges of applying ml to transcriptomic data, characterised by its remarkable diversity and complexity. Here, we introduce rnalib, a python library designed to facilitate the development of novel bioinformatics analysis methods for interpreting transcriptomics data. We present omicml, an intuitive graphical user interface (gui) that combines transcriptomic data analysis with machine learning (ml) based classification via integrating r and python packages libraries. Participants will learn how to import, clean, explore, and analyze genomics, transcriptomics, and proteomics data using widely adopted libraries and packages. The advent of pymportx extends far beyond mere data importation; it is about enabling researchers to harness the full power of python for transcriptomics analysis. The aim of this 3 day course is to empower researchers to start applying the fundamental scrna seq analysis pipeline to their own data. we will outline how to design and interpret results of a scrna seq dataset and, in teams, explore the basics of preprocessing and analysis in python on real data.
Comments are closed.