Transcript Table Ensembl Blog

Transcript Table Ensembl Blog Ensembl blog news about the ensembl project and its genome browser about us documentation projects. Transcript table the table shows all splice variants for a gene and includes noncoding transcripts. each transcript id includes a unique, stable 11 digit number. transcripts beginning with enst are human transcripts (for example, enst00000369985). a three letter code is inserted for other species; (for example, ensmust defines a mouse transcript).

Ensembl Genes And Transcripts Pdf Gene Genome Transcript table. at the top of the gene summary, the number of transcripts, or splice isoforms, are shown in a table. in our example, for ensembl release 114, there are nineteen transcripts for the human brca2 gene. you can find more information about the transcripts in the ‘biotype’ column. Each variation is a clickable link that will open a menu describing the genomic location, alleles, amino acid consequences, and links to get more information or see the variant in other transcript locations. there are more efficient ways to explore variants by gene, transcript and protein. It has three different sequence views, allowing you to view the exon and intron sequences in a table, an alignment of the cdna, cds and peptide sequences, and the protein sequence only. The ensembl genome browser provides a portal to sequence data, gene annotations predictions, and other types of data hosted in the various ensembl databases. many consider ensembl to be the most comprehensive and systematic gene annotation resource in the world.
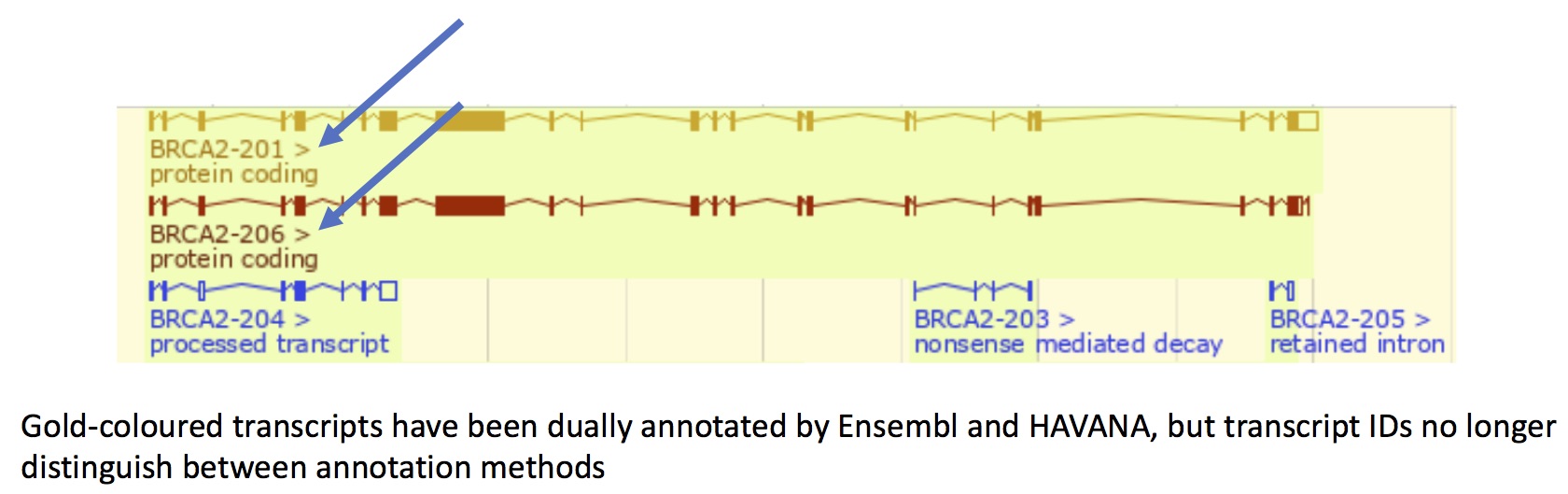
A Tweak To Ensembl Transcript Ids Ensembl Blog It has three different sequence views, allowing you to view the exon and intron sequences in a table, an alignment of the cdna, cds and peptide sequences, and the protein sequence only. The ensembl genome browser provides a portal to sequence data, gene annotations predictions, and other types of data hosted in the various ensembl databases. many consider ensembl to be the most comprehensive and systematic gene annotation resource in the world. For some species, the ensembl canonical transcript can be found in the tsv folder on the ensembl ftp site (at least it's there for humans in release 108). however, the file isn't there for all species, making it a bit difficult for us. Following this tutorial, i generated transcript counts with salmon, but now i'm stuck at the step of building a data frame transcripts to gene ids (tx2gene) using ensembldb package (for mouse). Gene constituents (transcript tab): return to the gene tab and show the transcript table. click on the id of the transcript selected in 2.2. to access the transcript tab. Introduction the ensembldb package provides functions to create and use transcript centric annotation databases packages. the annotation for the databases are directly fetched from ensembl 1 using their perl api.

Update To The Ensembl Canonical Transcript Set Ensembl Blog For some species, the ensembl canonical transcript can be found in the tsv folder on the ensembl ftp site (at least it's there for humans in release 108). however, the file isn't there for all species, making it a bit difficult for us. Following this tutorial, i generated transcript counts with salmon, but now i'm stuck at the step of building a data frame transcripts to gene ids (tx2gene) using ensembldb package (for mouse). Gene constituents (transcript tab): return to the gene tab and show the transcript table. click on the id of the transcript selected in 2.2. to access the transcript tab. Introduction the ensembldb package provides functions to create and use transcript centric annotation databases packages. the annotation for the databases are directly fetched from ensembl 1 using their perl api.
Comments are closed.