The Machinery Underlying A Molecular Dynamics Simulation

Molecular Dynamics Simulation Accuracy Speed Predictive Power This article explains how atomic motion is tracked by numerically solving equations of motion, the interatomic potential that is key to the simulation, and noteworthy machine learning force fields. you can learn about the fundamentals, applications, and latest technologies of molecular dynamics. This study comprehensively reviews the historical development, fundamental theories, and key techniques of md simulations, including all atom simulations, coarse grained simulations, and hybrid quantum mechanics molecular mechanics (qm mm) simulations.
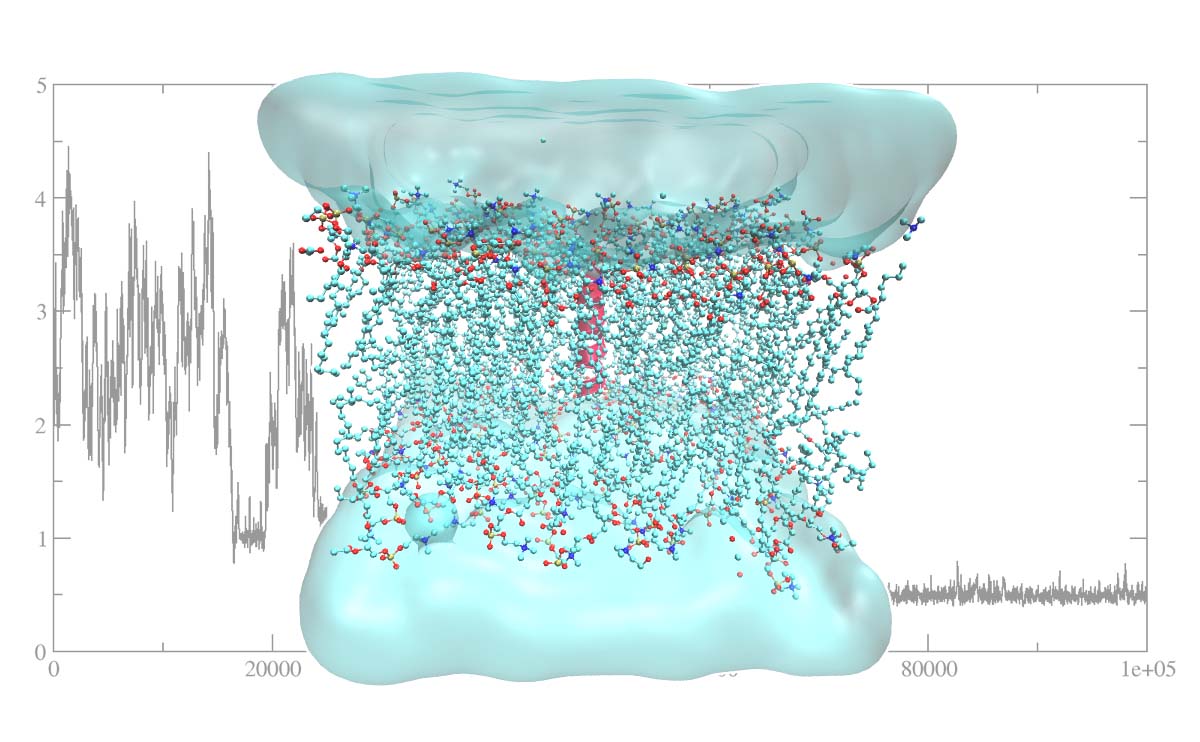
Getting Started With Molecular Dynamics Simulation Insilicosci The impact of molecular dynamics (md) simulations in molecular biology and drug discovery has expanded dramatically in recent years. these simulations capture the behavior of proteins and other biomolecules in full atomic detail and at very fine temporal resolution. In this approach, semi empirical models are used to simulate the interatomic interactions between constituent atoms. these semi empirical models define the interatomic forces, which are then used to calculate the potential energy of the material system under consideration. This chapter discusses a broad plethora of possibilities arising in drug discovery from molecular docking to md simulation with recent developments in these techniques and elaborates on the challenges and achievements of the focused approaches. In this tutorial, we present a cursory introduction to three widely used ml techniques─logistic regression, random forest, and multilayer perceptron─applied toward analyzing molecular dynamics (md) trajectory data.

Molecular Dynamics Simulation Photos Download The Best Free Molecular This chapter discusses a broad plethora of possibilities arising in drug discovery from molecular docking to md simulation with recent developments in these techniques and elaborates on the challenges and achievements of the focused approaches. In this tutorial, we present a cursory introduction to three widely used ml techniques─logistic regression, random forest, and multilayer perceptron─applied toward analyzing molecular dynamics (md) trajectory data. We used simulations to determine where this molecule binds to its receptor, and how it changes the binding strength of molecules that bind elsewhere (in part by changing the protein’s structure). We provide a general overview of the tools, force field, stages, applications, limitations, and future prospects of molecular dynamics in this paper. molecular dynamics is performed using tools such as gromacs, amber, charmm gui, namd, and lammps. Molecular dynamics (md) simulations employing classical force fields constitute the cornerstone of contemporary atomistic modeling in chemistry, biology, and materials science. however, the. Molecular dynamics (md) simulations rely heavily on force fields to model the interactions within and between molecules, providing a computational framework for studying molecular systems.

Molecular Dynamics Hardware Speed Accuracy Efficiency We used simulations to determine where this molecule binds to its receptor, and how it changes the binding strength of molecules that bind elsewhere (in part by changing the protein’s structure). We provide a general overview of the tools, force field, stages, applications, limitations, and future prospects of molecular dynamics in this paper. molecular dynamics is performed using tools such as gromacs, amber, charmm gui, namd, and lammps. Molecular dynamics (md) simulations employing classical force fields constitute the cornerstone of contemporary atomistic modeling in chemistry, biology, and materials science. however, the. Molecular dynamics (md) simulations rely heavily on force fields to model the interactions within and between molecules, providing a computational framework for studying molecular systems.

Molecular Dynamics Simulation Pipeline Download Scientific Diagram Molecular dynamics (md) simulations employing classical force fields constitute the cornerstone of contemporary atomistic modeling in chemistry, biology, and materials science. however, the. Molecular dynamics (md) simulations rely heavily on force fields to model the interactions within and between molecules, providing a computational framework for studying molecular systems.

Molecular Dynamics Simulation Model Download Scientific Diagram
Comments are closed.