Spls Da Comp1 2 Mixomics

Spls Da Comp1 2 Mixomics This page provides a quick start guide for applying partial least squares discriminant analysis (pls da) and its sparse variant (spls da) using mixomics. pls da is the special case of pls where the y dataframe is a single, categorical variable (y). In this practical we give an illustration of multivariate analysis for a supervised analysis context, but we will first start with a preliminary investigation with pca analysis on the gene expression data. the aim of this analysis is to select the genes that can help predict the class of the samples. how does this practical work?.

Multi Spls Da Comp1 2 Mixomics Splsda function fits an spls model with 1, … , ncomp components to the factor or class vector y. the appropriate indicator (dummy) matrix is created. An overview as part of mixomics: rohart f, gautier b, singh a, lê cao k a (2017) mixomics: an r package for ‘omics feature selection and multiple data integration. In the current version of mixomics, covariates that may confound the analysis are not included in the methods. we recommend correcting for those covariates beforehand using appropriate univariate or multivariate methods for batch effect removal. An overview as part of mixomics: rohart f, gautier b, singh a, l cao k a (2017) mixomics: an r package for omics feature selection and multiple data integration.
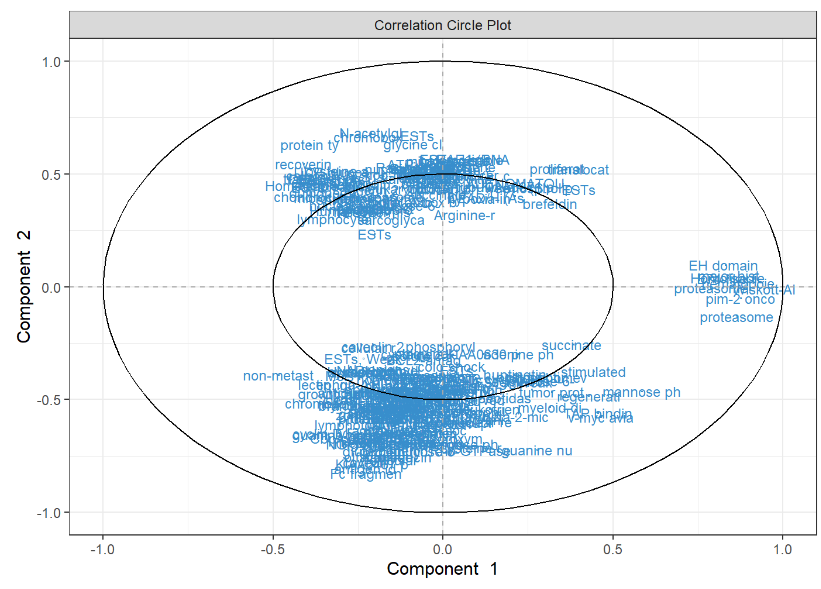
Spls Da Srbct Variable Plot Mixomics In the current version of mixomics, covariates that may confound the analysis are not included in the methods. we recommend correcting for those covariates beforehand using appropriate univariate or multivariate methods for batch effect removal. An overview as part of mixomics: rohart f, gautier b, singh a, l cao k a (2017) mixomics: an r package for omics feature selection and multiple data integration. Mixomics is an r toolkit dedicated to the exploration and integration of biological data sets with a specific focus on variable selection. the package currently includes more than twenty multivariate methodologies, mostly developed by the mixomics team (see some of our references in 1.2.3). Development repository for the bioconductor package 'mixomics ' mixomicsteam mixomics. The data are directly available in a processed and normalised format from the mixomics package. the small round blue cell tumours (srbct) dataset includes the expression levels of 2,308 genes measured on 63 samples. Integrate two datasets with pls da, a supervised approach to project individuals described by numeric variables (in a first dataset, x) on a subspace where they are separated at best based on their value for a categorical variable (in a second dataset, y).

Mixomics Omics Data Integration Project Mixomics is an r toolkit dedicated to the exploration and integration of biological data sets with a specific focus on variable selection. the package currently includes more than twenty multivariate methodologies, mostly developed by the mixomics team (see some of our references in 1.2.3). Development repository for the bioconductor package 'mixomics ' mixomicsteam mixomics. The data are directly available in a processed and normalised format from the mixomics package. the small round blue cell tumours (srbct) dataset includes the expression levels of 2,308 genes measured on 63 samples. Integrate two datasets with pls da, a supervised approach to project individuals described by numeric variables (in a first dataset, x) on a subspace where they are separated at best based on their value for a categorical variable (in a second dataset, y).
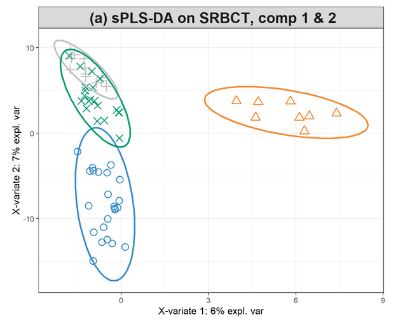
Webinars Mixomics The data are directly available in a processed and normalised format from the mixomics package. the small round blue cell tumours (srbct) dataset includes the expression levels of 2,308 genes measured on 63 samples. Integrate two datasets with pls da, a supervised approach to project individuals described by numeric variables (in a first dataset, x) on a subspace where they are separated at best based on their value for a categorical variable (in a second dataset, y).

Rplot08 Mixomics
Comments are closed.