Space Efficient Sequence Alignment

Sequence Alignment Methods And Algorithms Pdf Sequence Alignment The sequence alignment problem is fundamental in bioinformatics; we have implemented the x drop algorithm, a heuristic method for pairwise alignment that reduces search space, on the graphcore intelligence processor unit (ipu) accelerator. The sequence alignment problem is fundamental in bioinformatics; we have implemented the $x$ drop algorithm, a heuristic method for pairwise alignment that reduces search space, on the.

Lecture 6 Evolutionary Sequence Alignment Algorithms Pdf Sequence In this review, pairwise sequence alignment and its scoring system, main algorithms for multiple sequence alignment, as well as their advantages and disadvantages, and the quality estimation methods for multiple sequence alignment software, are presented and discussed. Good filters reduce the search space before alignment without missing significant matches. we introduce dream stellar, a parallelized, updated version of the pairwise local aligner stellar. Alignment score • space complexity of computing just the score itself is o(n) we only need the previous column to calculate the current column, and we can then throw away that previous column once we’re done using it. We develop a space efficient algorithm to compute such an optimal alignment which consumes only o (nm) space within the same amount of time. our algorithm enables the computation for a pair of dna sequences of length up to 10,000 to be carried out on an ordinary desktop computer.
Github Jiazheng Yuan Sequence Alignment Space Efficient Iterative Alignment score • space complexity of computing just the score itself is o(n) we only need the previous column to calculate the current column, and we can then throw away that previous column once we’re done using it. We develop a space efficient algorithm to compute such an optimal alignment which consumes only o (nm) space within the same amount of time. our algorithm enables the computation for a pair of dna sequences of length up to 10,000 to be carried out on an ordinary desktop computer. Learn how to perform sequence alignments using linear space algorithms to efficiently compare biological sequences with reduced memory usage. Dedicated accelerator hardware has become essential for processing ai based workloads, leading to the rise of novel accelerator architectures. furthermore, fund. The sequence alignment problem is fundamental in bioinformatics; we have implemented the x drop algorithm, a heuristic method for pairwise alignment that reduces search space, on the graphcore intelligence processor unit (ipu) accelerator. We introduce lexicmap, a nucleotide sequence alignment tool for efficiently querying moderate length sequences (>250 bp) such as a gene, plasmid or long read against up to millions of.

Sequence Alignment Assignment Point Learn how to perform sequence alignments using linear space algorithms to efficiently compare biological sequences with reduced memory usage. Dedicated accelerator hardware has become essential for processing ai based workloads, leading to the rise of novel accelerator architectures. furthermore, fund. The sequence alignment problem is fundamental in bioinformatics; we have implemented the x drop algorithm, a heuristic method for pairwise alignment that reduces search space, on the graphcore intelligence processor unit (ipu) accelerator. We introduce lexicmap, a nucleotide sequence alignment tool for efficiently querying moderate length sequences (>250 bp) such as a gene, plasmid or long read against up to millions of.
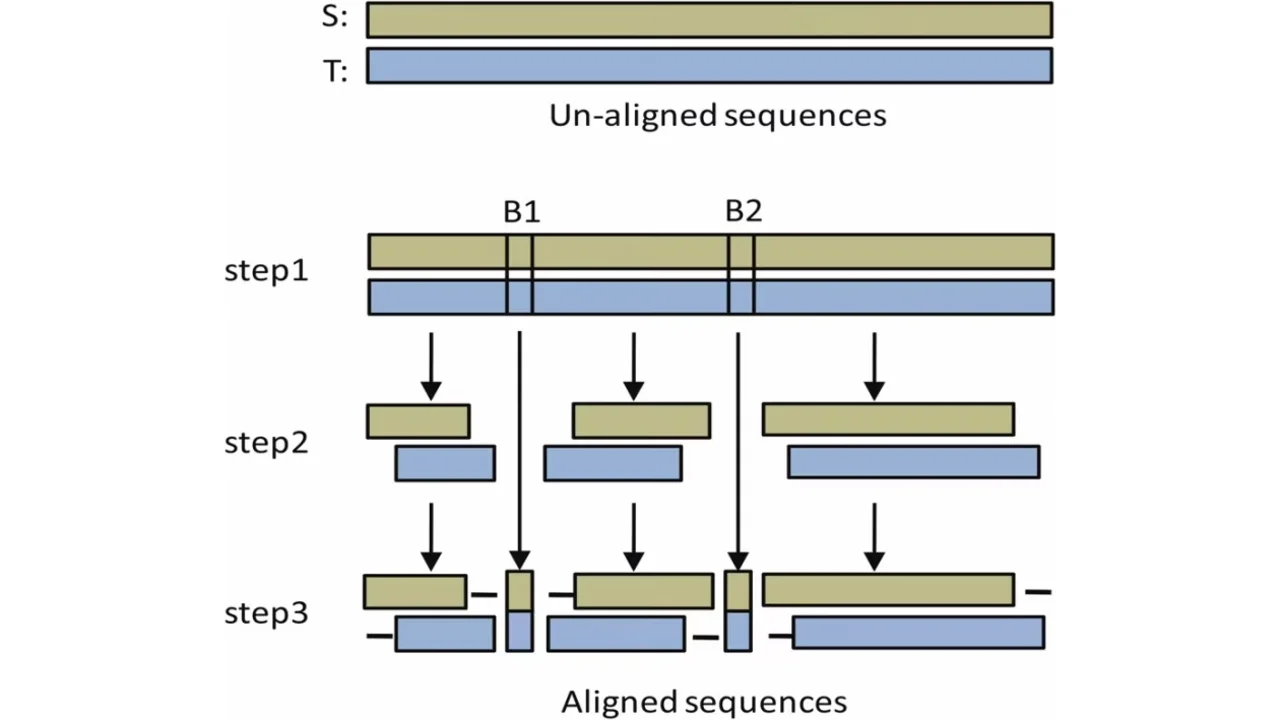
Efficient Sequence Alignment Technique For Bioinformatics Sensors The sequence alignment problem is fundamental in bioinformatics; we have implemented the x drop algorithm, a heuristic method for pairwise alignment that reduces search space, on the graphcore intelligence processor unit (ipu) accelerator. We introduce lexicmap, a nucleotide sequence alignment tool for efficiently querying moderate length sequences (>250 bp) such as a gene, plasmid or long read against up to millions of.
Comments are closed.