Solved Second Generation Sequencing Differs From Sanger Chegg

Solved Second Generation Sequencing Differs From Sanger Chegg Second generation sequencing does not depend on chain terminator nucleoside triphosphates. second generation sequencing does not require dna polymerase. for the cost per base sequenced, second generation sequencing is much more expensive than sanger sequencing. here’s the best way to solve it. In summary, the key aspect that sets second generation sequencing apart from sanger sequencing is its ability to process a large volume of sequences simultaneously, significantly reducing time and costs while increasing throughput for complex genomic analyses.
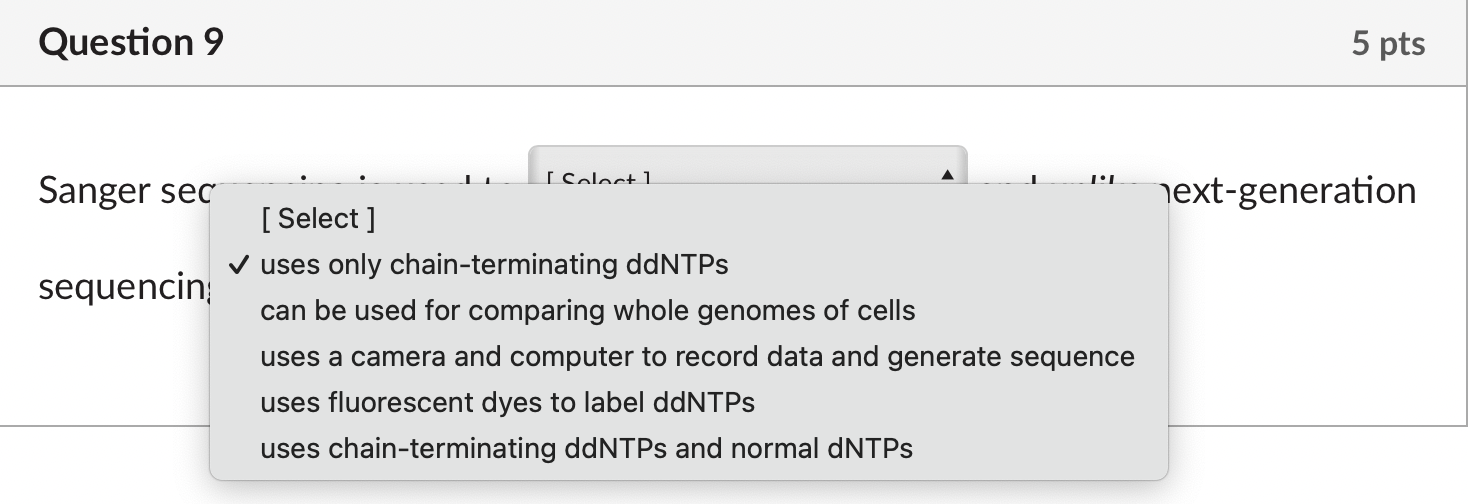
Solved Sanger Sequencing Is Used To And Unlike Chegg In summary, first generation sequencing (sanger sequencing) offers high accuracy and long read lengths but is low throughput and costly. second generation sequencing (ngs) provides high throughput and cost effectiveness but has shorter read lengths and higher error rates. Second and third generation sequencing methods have been developed with improvements in cost, time, and accuracy of sequencing. the data obtained from these methods are interpreted with bioinformatics and contributed to the development of next generation sequencing technology. Sanger is traditional and low throughput; ngs is modern and high throughput. low throughput; sequences one fragment at a time (up to ~1 kb). high throughput; can sequence millions of fragments in parallel. ngs is better for large scale projects like whole genome sequencing. long reads (~500–1000 bp). Wondering about the difference between next gen and sanger sequencing? this article explores technical and applied differences in the technologies.
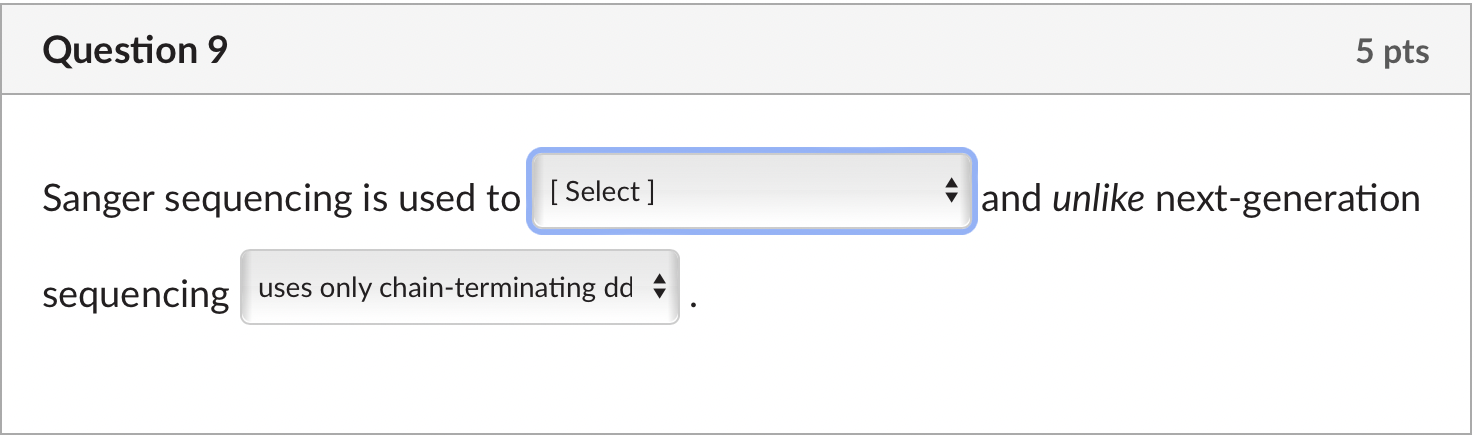
Solved Sanger Sequencing Is Used To And Unlike Chegg Sanger is traditional and low throughput; ngs is modern and high throughput. low throughput; sequences one fragment at a time (up to ~1 kb). high throughput; can sequence millions of fragments in parallel. ngs is better for large scale projects like whole genome sequencing. long reads (~500–1000 bp). Wondering about the difference between next gen and sanger sequencing? this article explores technical and applied differences in the technologies. Nanopore sequencing involves passing a dna molecule through a tiny pore and measuring the change in electrical current to determine the sequence, while sanger sequencing uses dideoxynucleotides to terminate dna synthesis at different points, allowing the sequence to be determined. Compare sanger sequencing vs. ngs, exploring key differences, advantages, and applications in modern dna sequencing techniques. Next generation sequencing (ngs) and sanger sequencing are two widely used methods for determining the nucleotide sequence of dna. while both techniques are valuable tools in molecular biology, they have distinct differences in terms of speed, cost, accuracy, and applications. Students usually get confused between these terms (first, second, third and next generation sequencing). in this article, i will explain to you sequencing generations, their advantages, limitations and applications.
Comments are closed.