Sequence Alignment Exercise Gt Computability Complexity Theory Algorithms

Comparing Runtime Efficiency Of Sequence Alignment Algorithms In Pytho Sequence alignment exercise gt computability, complexity, theory: algorithms udacity 646k subscribers subscribed. Sequence alignment exercise gt computability complexity theory algorithms lesson with certificate for programming courses.

Ppt Sample Complexity Of Algorithm Configuration For Sequence We concatenate the two alignments obtained for every optimal point recursively and in turn find the optimal score for the original two input strings as well as the final alignment. the memory efficient version is implemented in o (m n) space. An alignment is an assignment of gaps to positions 0,…, n in x, and 0,…, n in y, so as to line up each letter in one sequence with either a letter, or a gap in the other sequence. An alignment m is a set of ordered pairs xi yj such that each item occurs in at most one pair and no crossings. def. the pair xi yj and xi' yj' cross if i < i', but j > j'. q. ctaccg vs. tacatg. suppose all α=1 for all mismatches and δ=1, what then is cost of m = {x2 y1, x3 y2, x4 y3, x5 y4, x6 y6} ? q. ctaccg vs. tacatg. The sequence alignment problem takes as input two or more sequences, and produces as output an arrangement of those sequences that highlights their similarities and differences.

Exercises Download Free Pdf Computational Complexity Theory An alignment m is a set of ordered pairs xi yj such that each item occurs in at most one pair and no crossings. def. the pair xi yj and xi' yj' cross if i < i', but j > j'. q. ctaccg vs. tacatg. suppose all α=1 for all mismatches and δ=1, what then is cost of m = {x2 y1, x3 y2, x4 y3, x5 y4, x6 y6} ? q. ctaccg vs. tacatg. The sequence alignment problem takes as input two or more sequences, and produces as output an arrangement of those sequences that highlights their similarities and differences. Use alignment algorithms to find optimal alignment. different alignment algorithms (global, local, affine penalty). types of two sequence alignments: input: treat the two sequences as potentially equivalent. goal: identify conserved regions and differences. Ve s and a c st n s and ends at vi. describe a dynamic programming algorithm that computes ts;i fo a em s or exactly 2 literals. the point of this problem is to show that the satis ability problem for formulas in 2cnf can be solved by a polyno n exactly two ways. for instance (p :q) i equivalent to (q ! p) and (:p ! :q), and (p q) is. In this review, pairwise sequence alignment and its scoring system, main algorithms for multiple sequence alignment, as well as their advantages and disadvantages, and the quality estimation methods for multiple sequence alignment software, are presented and discussed. Local alignment motivation useful for comparing protein sequences that share a common motif (conserved pattern) or domain (independently folded unit) but differ elsewhere.
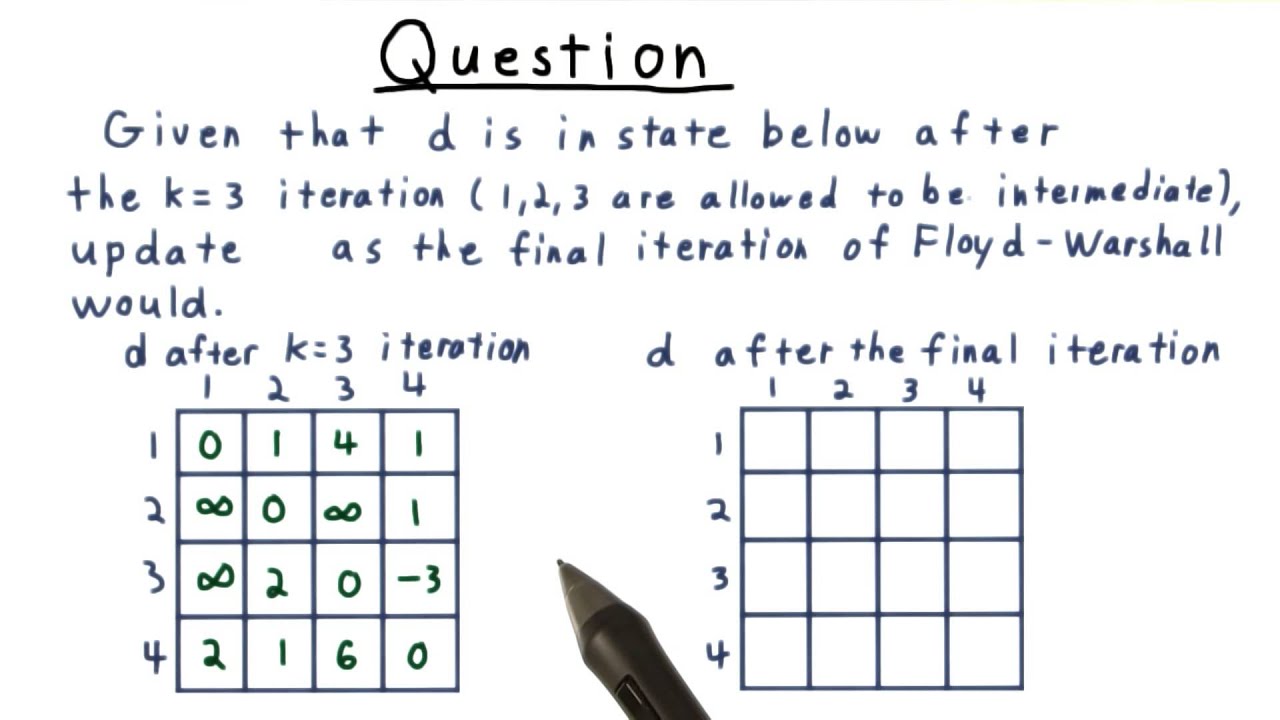

Floyd Warshall Algorithm Exercise Gt Computability Complexity Use alignment algorithms to find optimal alignment. different alignment algorithms (global, local, affine penalty). types of two sequence alignments: input: treat the two sequences as potentially equivalent. goal: identify conserved regions and differences. Ve s and a c st n s and ends at vi. describe a dynamic programming algorithm that computes ts;i fo a em s or exactly 2 literals. the point of this problem is to show that the satis ability problem for formulas in 2cnf can be solved by a polyno n exactly two ways. for instance (p :q) i equivalent to (q ! p) and (:p ! :q), and (p q) is. In this review, pairwise sequence alignment and its scoring system, main algorithms for multiple sequence alignment, as well as their advantages and disadvantages, and the quality estimation methods for multiple sequence alignment software, are presented and discussed. Local alignment motivation useful for comparing protein sequences that share a common motif (conserved pattern) or domain (independently folded unit) but differ elsewhere.
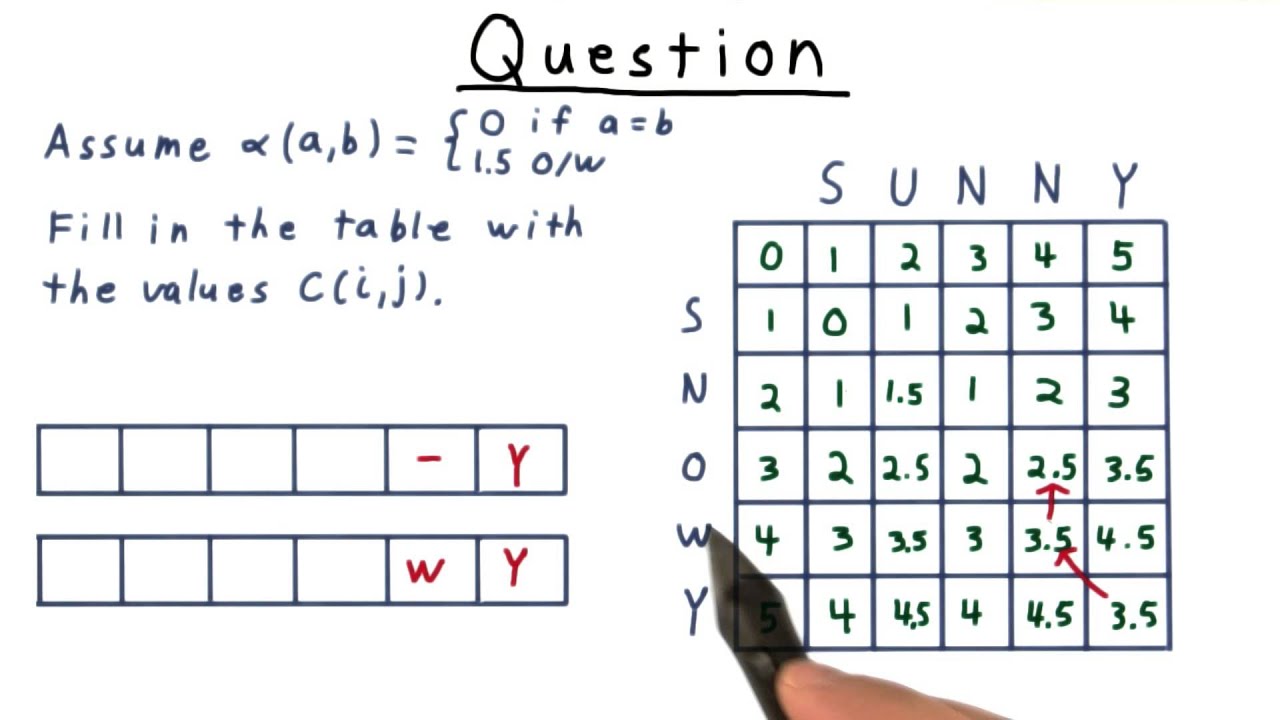
Sequence Alignment Solution Gt Computability Complexity Theory In this review, pairwise sequence alignment and its scoring system, main algorithms for multiple sequence alignment, as well as their advantages and disadvantages, and the quality estimation methods for multiple sequence alignment software, are presented and discussed. Local alignment motivation useful for comparing protein sequences that share a common motif (conserved pattern) or domain (independently folded unit) but differ elsewhere.
Comments are closed.