Scpubr Do Nebulosaplot

Scpubr Do nebulosaplot: wrapper for nebulosa::plot density in seurat. description wrapper for nebulosa::plot density in seurat. usage do nebulosaplot( sample, features, slot = null, dims = c(1, 2), pt.size = 1, reduction = null, combine = true, method = c("ks", "wkde"), joint = false, return only joint = false, plot.title = null, plot.subtitle = null. Nebulosa plot with scpubr. then, this type visualization becomes a natural partner to `seurat::featureplot ()’ as not only we are able to visualize the expression of a variable, but also query the density of the surrounding cells. here is an example: comparison of a nebulosa vs a featureplot.
Github Enblacar Scpubr Generate High Quality Publication Ready In scpubr: generate publication ready visualizations of single cell transcriptomics data view source: r do nebulosaplot.r. Here, we perform an integrative single cell and spatial transcriptomic analysis on hpv negative oral squamous cell carcinoma (oscc) to comprehensively characterize malignant cells in tumor core. Scpubr provides a streamlined way of generating publication ready plots for known s ingle c ell visualizations in a " pub lication r eady" format (scpubr). this is, the aim is to automatically generate plots with the highest quality possible, that can be used right away or with minimal modifications for a research article. Do groupwisedeplot compute a heatmap with the results of a group wise de analysis.
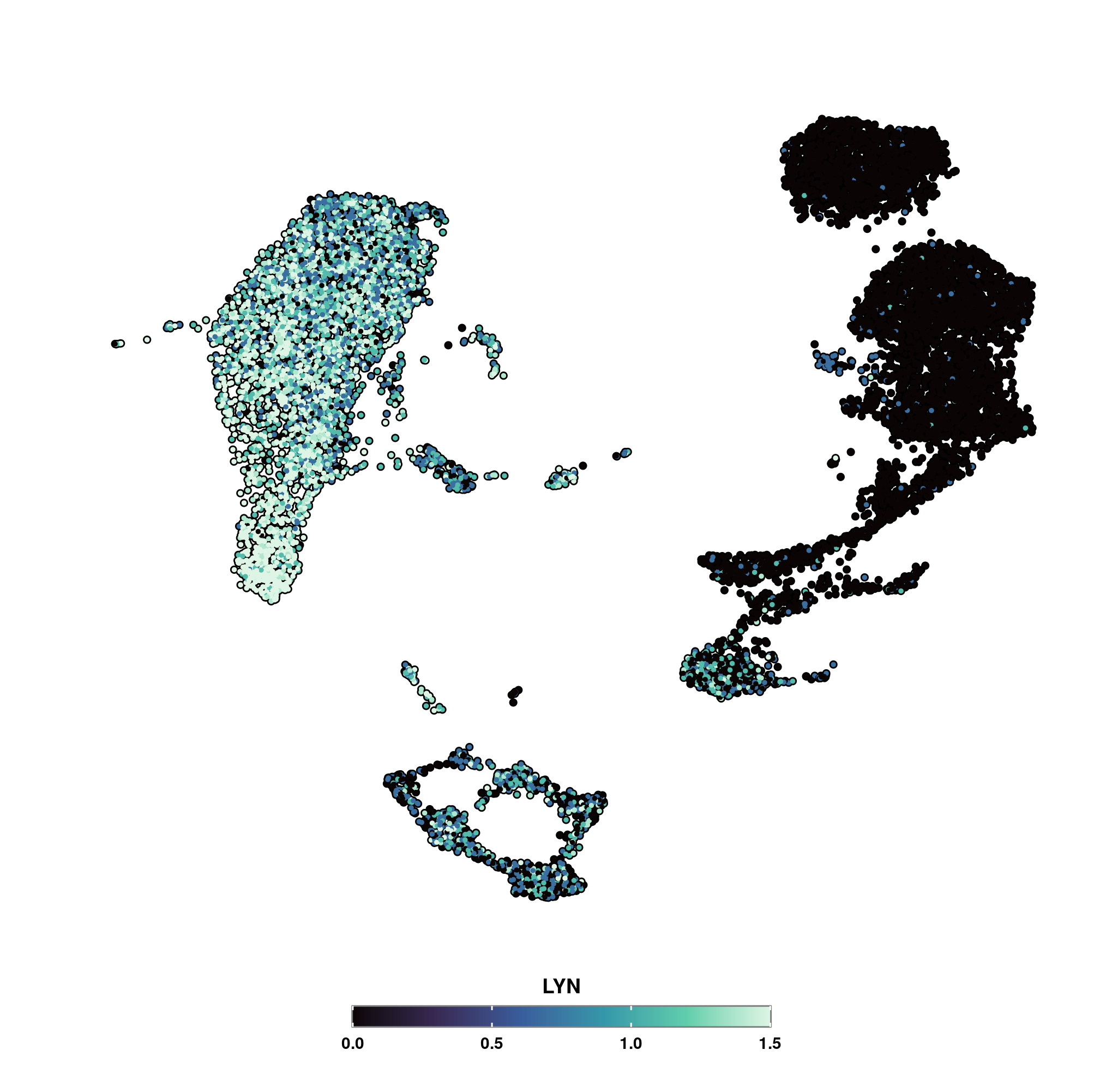
Feature Request Add Min Max Cutoff Do Featureplot Issue 2 Scpubr provides a streamlined way of generating publication ready plots for known s ingle c ell visualizations in a " pub lication r eady" format (scpubr). this is, the aim is to automatically generate plots with the highest quality possible, that can be used right away or with minimal modifications for a research article. Do groupwisedeplot compute a heatmap with the results of a group wise de analysis. Wrapper for nebulosa::plot density in seurat. sample, features, slot = null, dims = c(1, 2), pt.size = 1, reduction = null, combine = true, method = c("ks", "wkde"), joint = false, return only joint = false, plot.title = null, plot.subtitle = null, plot.caption = null, legend.type = "colorbar", legend.framewidth = 0.5, legend.tickwidth = 0.5,. Description a system that provides a streamlined way of generating publication ready plots for known single cell transcriptomics data in a “publication ready” format. this is, the goal is to automatically generate plots with the highest quality possible, that can be used right away or with minimal modifications for a research article. Do nebulosaplot () | density based feature visualization nebulosa plots use kernel density estimation to visualize where cells expressing a feature are concentrated in the dimensional reduction space. Any scripts or data that you put into this service are public.

Feature Request Add Min Max Cutoff Do Featureplot Issue 2 Wrapper for nebulosa::plot density in seurat. sample, features, slot = null, dims = c(1, 2), pt.size = 1, reduction = null, combine = true, method = c("ks", "wkde"), joint = false, return only joint = false, plot.title = null, plot.subtitle = null, plot.caption = null, legend.type = "colorbar", legend.framewidth = 0.5, legend.tickwidth = 0.5,. Description a system that provides a streamlined way of generating publication ready plots for known single cell transcriptomics data in a “publication ready” format. this is, the goal is to automatically generate plots with the highest quality possible, that can be used right away or with minimal modifications for a research article. Do nebulosaplot () | density based feature visualization nebulosa plots use kernel density estimation to visualize where cells expressing a feature are concentrated in the dimensional reduction space. Any scripts or data that you put into this service are public.
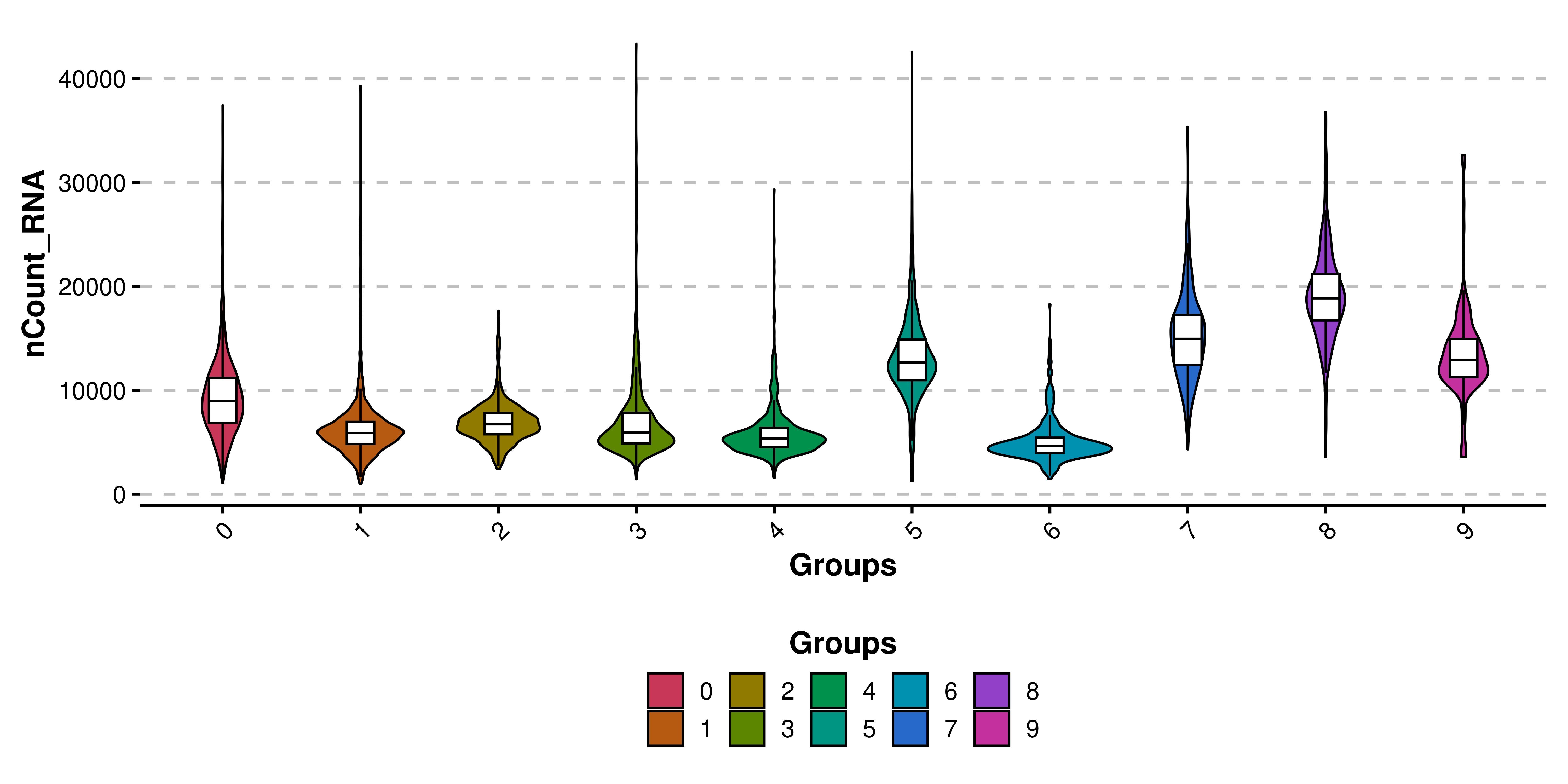
Scpubr Do Beeswarmplot Do nebulosaplot () | density based feature visualization nebulosa plots use kernel density estimation to visualize where cells expressing a feature are concentrated in the dimensional reduction space. Any scripts or data that you put into this service are public.
Comments are closed.