Scalable Spatial Transcriptomics Through Computational Array

Computational Approaches And Challenges In Spatial Transcriptomics Here, we present an imaging free spatial transcriptomics method that computationally reconstructs array barcode locations used in spatial transcriptomics measurements with high resolution. Here, we present an imaging free spatial transcriptomics method that computationally reconstructs array barcode locations used in spatial transcriptomics measurements with high resolution and fidelity.
Scalable Spatial Transcriptomics Via Computational Array Reconstruction Spatial transcriptomics enables gene expression mapping within tissues but is often limited by imaging constraints. we present an imaging free approach that reconstructs spatial barcode. Published in nature biotechnology, their work describes a computational framework that reconstructs the spatial locations of transcriptomic barcodes through an innovative approach leveraging molecular diffusion patterns paired with dimensionality reduction algorithms. Spatial transcriptomics enables gene expression mapping within tissues but is often limited by imaging constraints. we present an imaging free approach that reconstructs spatial barcode locations using molecular diffusion and dimensionality reduction. These approaches leads to a challenging computational array reconstruction problem. we show that a statistical inference method, which we term accumap, outperforms the current approach which is to run umap on the bead count matrix.

Updated Computational Tools For Spatial Transcriptomics Analysis Spatial transcriptomics enables gene expression mapping within tissues but is often limited by imaging constraints. we present an imaging free approach that reconstructs spatial barcode locations using molecular diffusion and dimensionality reduction. These approaches leads to a challenging computational array reconstruction problem. we show that a statistical inference method, which we term accumap, outperforms the current approach which is to run umap on the bead count matrix. Scalable spatial transcriptomics through computational array reconstruction by jeremy charette | april 3, 2025 | comments off. Both the methodological and computational solutions for spatial transcriptomics are evolving at a fast pace, providing coverage to larger sample areas at higher resolution thus producing spatial datasets with increasing complexity and size.

Updated Computational Tools For Spatial Transcriptomics Analysis Scalable spatial transcriptomics through computational array reconstruction by jeremy charette | april 3, 2025 | comments off. Both the methodological and computational solutions for spatial transcriptomics are evolving at a fast pace, providing coverage to larger sample areas at higher resolution thus producing spatial datasets with increasing complexity and size.

Scalable Spatial Transcriptomics Through Computational Array
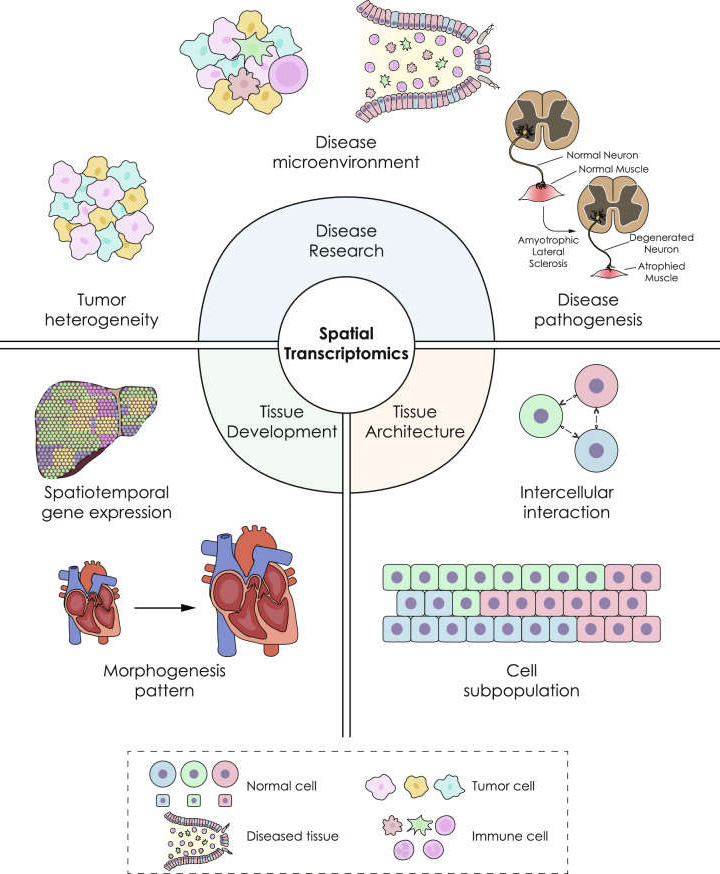
Spatial Transcriptomics Solutions Creative Bioarray Creative Bioarray
Comments are closed.