Sample Level Enrichment Analysis Slea
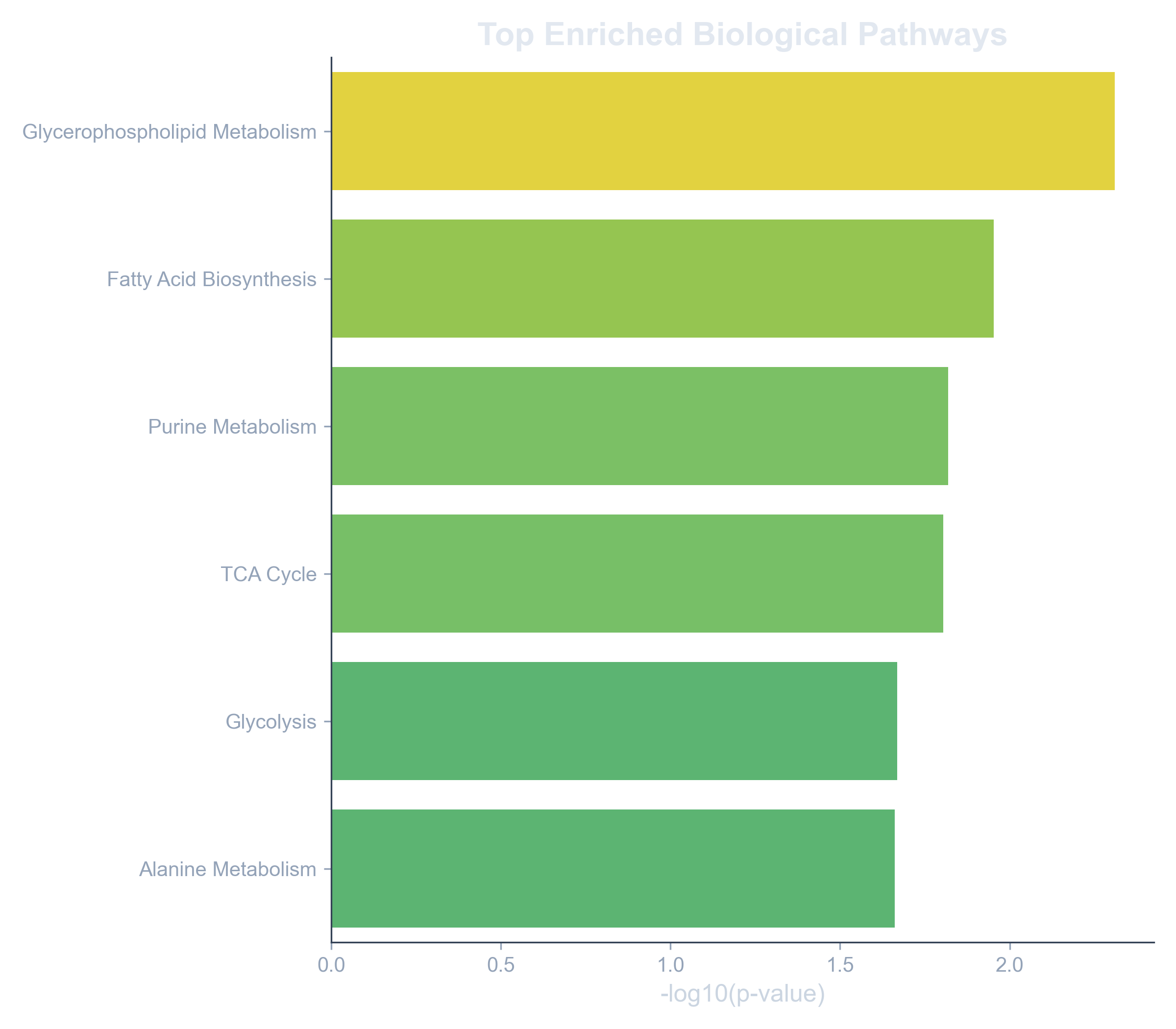
Enrichment Analysis Metabonexus Slea is a method to analyse the transcriptional status of gene modules (gene sets) per sample in a transcriptomic dataset. the results are presented in the form of interactive heatmaps which facilitates their interpretation. With this analysis we can asses the transcriptional status of each pathway (or other modules) for each sample. in a second step we can relate the enrichment status with the clinical annotation, like cancer subtypes. to see how it is done, watch the video tutorial and or read the instructions below.

Slea Sample Level Enrichment Analysis In this manuscript, we propose a new methodology, sample level enrichment analysis (slea), that overcomes these limitations and has a more general use for enrichment analysis (ea) at the level of samples. We propose a new approach based on enrichment analysis at the level of samples (sample level enrichment analysis slea) in expression profiling datasets. without using a priori. In this video it is shown how with the help of gitools you can analyze the transcription level status of different pathways and then view it in context of clinical data .more. This approach provides insights into the collective behavior of functionally related genes, revealing potential biological processes associated with the observed changes in gene expression. enrichment analysis involves several steps that we will look at individually in the following sections.

Slea Sample Level Enrichment Analysis In this video it is shown how with the help of gitools you can analyze the transcription level status of different pathways and then view it in context of clinical data .more. This approach provides insights into the collective behavior of functionally related genes, revealing potential biological processes associated with the observed changes in gene expression. enrichment analysis involves several steps that we will look at individually in the following sections. We provide an overview of existing methods for enrichment analysis of a given gene list data with regard to functional gene sets and pathways. functionality for differential expression analysis, set and network based enrichment analysis, along with visualization and exploration of results. Without using a priori phenotypic information about samples, slea calculates an enrichment score per sample per gene set using z test. Without using a priori phenotypic information about samples, slea calculates an enrichment score per sample per gene set using z test.

Slea Sample Level Enrichment Analysis We provide an overview of existing methods for enrichment analysis of a given gene list data with regard to functional gene sets and pathways. functionality for differential expression analysis, set and network based enrichment analysis, along with visualization and exploration of results. Without using a priori phenotypic information about samples, slea calculates an enrichment score per sample per gene set using z test. Without using a priori phenotypic information about samples, slea calculates an enrichment score per sample per gene set using z test.
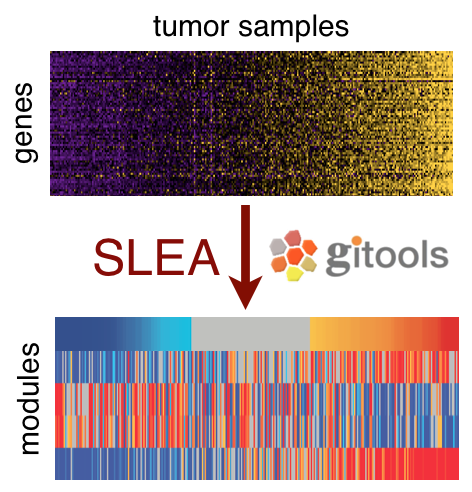
Slea Sample Level Enrichment Analysis Without using a priori phenotypic information about samples, slea calculates an enrichment score per sample per gene set using z test.

Functional Enrichment Analysis A C The Gene Set Enrichment Analysis
Comments are closed.