Protocol Visualizing Gene Expression Method

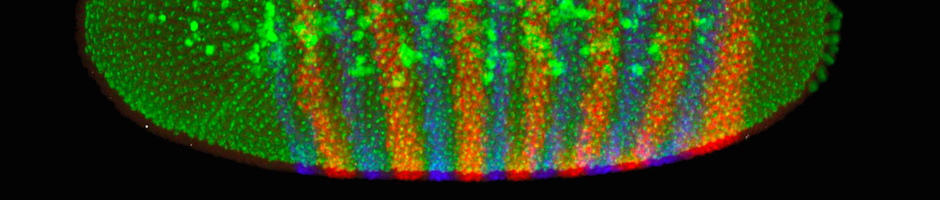
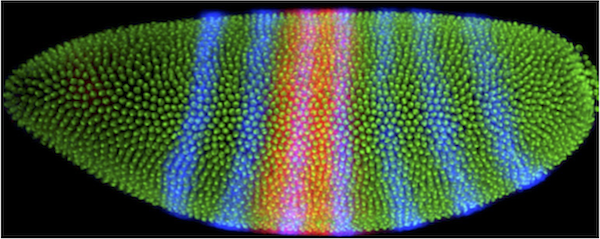
Protocol Visualizing Gene Expression Method To study how different cell types emerge during development through the action of different dna sequences, we visualize mrnas in fruit fly embryos. we study fruit flies because they have a surprising amount in common with humans and are easier and more ethical to work with. With this visualization we can appreciate the spatial localization of gene expression changes and observe the emergence of downregulated genes at 30 min, with robust negative fold changes in mrnas encoding for proteins localized in the endoplasmic reticulum.
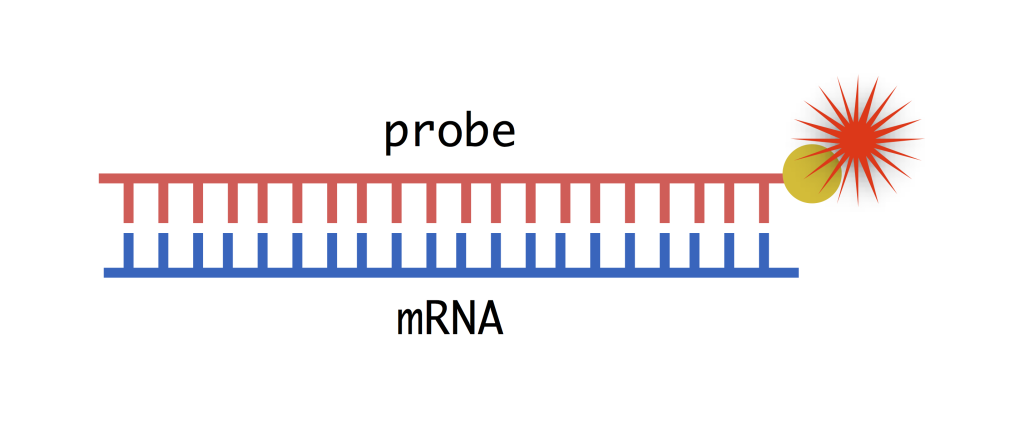
Protocol Visualizing Gene Expression Method We show that this multimodal approach is applicable to gene expression visualized by protein specific antibodies and fluorescence rna in situ hybridisation offering a detailed understanding. First, we will describe fluorescence imaging of gene expression with a wide variety of fluorescent proteins. next, we will discuss the strategies for bli of gene expression. besides incorporating the reporter gene into the host dna, mrna based bli of gene expression will also be briefly mentioned. Cutting edge and thorough, imaging gene expression: methods and protocols, second edition is a valuable resource for any researcher interested in learning more about this evolving and important field. The tool allows users to upload gene expression datasets and interact with a dashboard to explore gene expression binarization and the inferred boolean networks.

Protocol Visualizing Gene Expression Method Cutting edge and thorough, imaging gene expression: methods and protocols, second edition is a valuable resource for any researcher interested in learning more about this evolving and important field. The tool allows users to upload gene expression datasets and interact with a dashboard to explore gene expression binarization and the inferred boolean networks. This protocol demonstrates how to perform integration of visium spatial gene expression data with single cell rna seq data using two tools: seurat 2 and giotto 3. Summary the last decade has witnessed massive advancements in high throughput techniques capable of producing increasingly complex gene expression datasets across time and space and at the resolution of single cells. In this review, we explain three main technologies: microarrays, rna sequencing (rna seq), and quantitative real time pcr (qpcr). we describe how each method works, their strengths and limitations, and the situations where they are best used. Explore advanced imaging techniques for real time gene expression monitoring in biotechnology. discover their applications and impact on research.
Protocol Visualizing Gene Expression Method This protocol demonstrates how to perform integration of visium spatial gene expression data with single cell rna seq data using two tools: seurat 2 and giotto 3. Summary the last decade has witnessed massive advancements in high throughput techniques capable of producing increasingly complex gene expression datasets across time and space and at the resolution of single cells. In this review, we explain three main technologies: microarrays, rna sequencing (rna seq), and quantitative real time pcr (qpcr). we describe how each method works, their strengths and limitations, and the situations where they are best used. Explore advanced imaging techniques for real time gene expression monitoring in biotechnology. discover their applications and impact on research.

Protocol Visualizing Gene Expression Method In this review, we explain three main technologies: microarrays, rna sequencing (rna seq), and quantitative real time pcr (qpcr). we describe how each method works, their strengths and limitations, and the situations where they are best used. Explore advanced imaging techniques for real time gene expression monitoring in biotechnology. discover their applications and impact on research.
Comments are closed.