Protein Folding Portfolio

Protein Folding Activity Pitcha Portfolio Here, the authors use ai based protein design software to uncover a broad range of protein folds that can be formed from small, primordial amino acid libraries constrained by asteroid and. Uncover the latest and most impactful research in protein folding. explore pioneering discoveries, insightful ideas and new methods from leading researchers in the field.

Protein Folding Activity Pitcha Portfolio In this work, we propose simplefold, a flow matching based folding model that directly maps a protein sequence to its full 3d atomic structure without relying on msa, pairwise interaction maps, triangular updates or any other equivariant geometric modules. We scale simplefold to 3b parameters and train it on more than 8.6m distilled protein structures together with experimental pdb data. to the best of our knowledge, simplefold is the largest scale folding model ever developed. Here, we review the implications of alphafold for protein folding, including the original questions about structure prediction and folding mechanism as well as the broader question about energy landscapes. The goal of this project is to find the best way to fold proteins using a hydrophobic polar protein folding model. the concept of the “best way” refers to a method for transforming a string into 2d or 3d shapes that best satisfy the criteria of hydrophobic polar protein folding.

Protein Folding Activity Pitcha Portfolio Here, we review the implications of alphafold for protein folding, including the original questions about structure prediction and folding mechanism as well as the broader question about energy landscapes. The goal of this project is to find the best way to fold proteins using a hydrophobic polar protein folding model. the concept of the “best way” refers to a method for transforming a string into 2d or 3d shapes that best satisfy the criteria of hydrophobic polar protein folding. We conduct comprehensive ablation studies to reveal the role of different types of protein features and model designs, inspiring further simplification and improvement. A number of active projects on folding@home right now aim to understand how different forms of the protein apolipoprotein e (apoe) determine one’s risk of developing alzheimer’s disease. Machine learning has revolutionized protein structure prediction and design. this review discusses current methods for protein folding and inverse folding challenges. While inferences about protein folding can be made through mutation studies, typically, experimental techniques for studying protein folding rely on the gradual unfolding or folding of proteins and observing conformational changes using standard non crystallographic techniques.

Folding Portfolio We conduct comprehensive ablation studies to reveal the role of different types of protein features and model designs, inspiring further simplification and improvement. A number of active projects on folding@home right now aim to understand how different forms of the protein apolipoprotein e (apoe) determine one’s risk of developing alzheimer’s disease. Machine learning has revolutionized protein structure prediction and design. this review discusses current methods for protein folding and inverse folding challenges. While inferences about protein folding can be made through mutation studies, typically, experimental techniques for studying protein folding rely on the gradual unfolding or folding of proteins and observing conformational changes using standard non crystallographic techniques.

Protein Folding Project E Portfolio Machine learning has revolutionized protein structure prediction and design. this review discusses current methods for protein folding and inverse folding challenges. While inferences about protein folding can be made through mutation studies, typically, experimental techniques for studying protein folding rely on the gradual unfolding or folding of proteins and observing conformational changes using standard non crystallographic techniques.
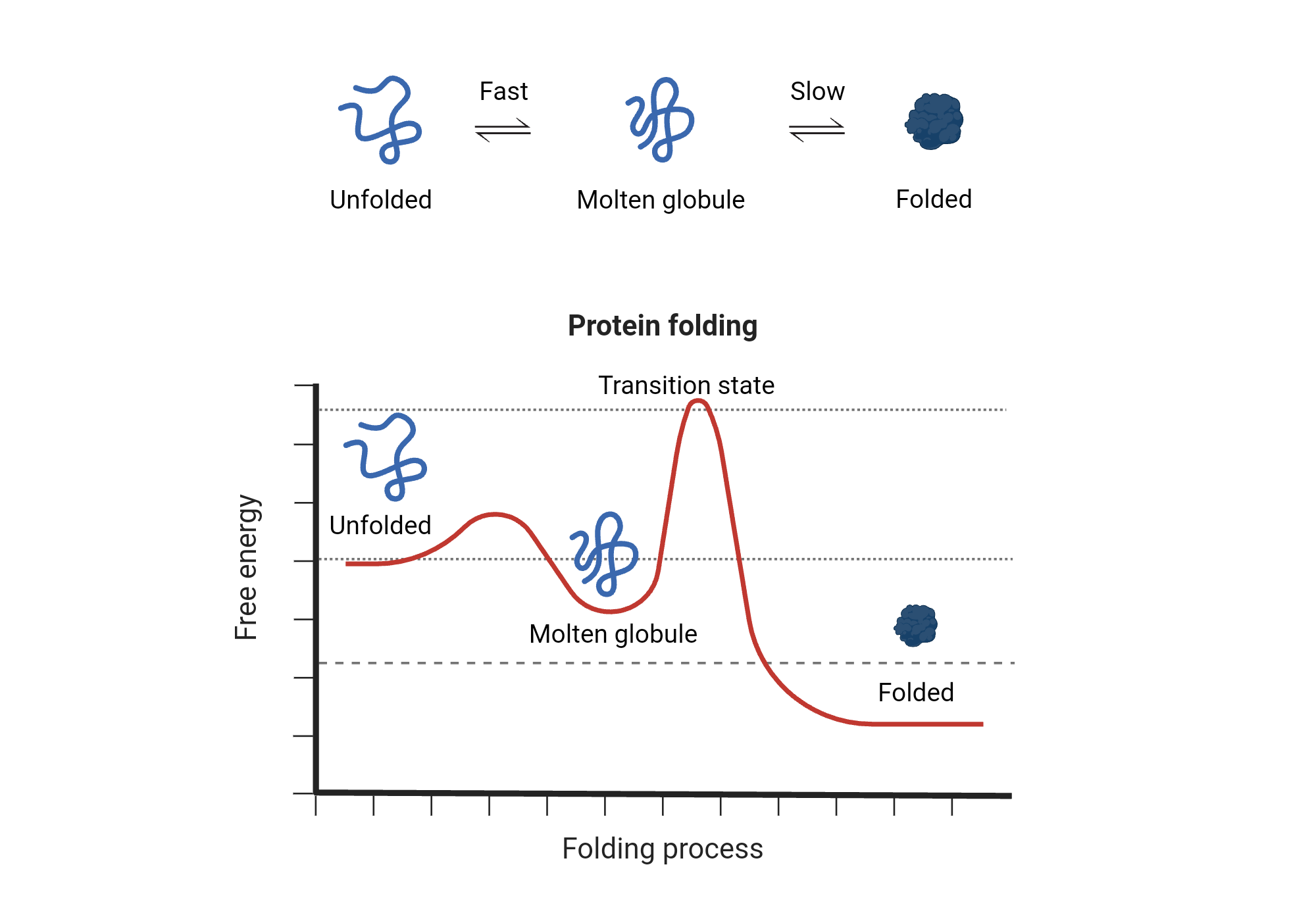
Protein Folding Biorender Science Templates
Comments are closed.