Pdf An Enhanced Web Based Tools For Multiple Sequence Alignment A

Tools For Multiple Sequence Alignment Department Of Bioinformatics This review provides an overview on the development of multiple sequence alignment (msa) methods and their main applications. it is focused on progress made over the past decade. Multiple sequence alignment (msa) is one of the oldest dilemmas in computational biology, involving the alignment of more than two dna, rna, or protein sequence.

Pdf An Enhanced Web Based Tools For Multiple Sequence Alignment A This paper presents probcons, a practical tool for progressive protein multiple sequence alignment based on probabilistic consistency, and evaluates its performance on several standard alignment benchmark data sets. In this paper, we present our new progressive alignment algorithm, called motalign, for multiple sequence alignment (msa). our algorithm adopts a new profile based distance [1] to compare two biological sequences and a new score function, called gpsp, to align profiles. we benchmarked our algorithm on different benchmarks. In this review, the pairwise sequence alignment algorithms and the corresponding scoring system, heuristic algorithms for multiple sequence alignment and their defects, and quality estimation methods used to test multiple sequence alignment software are reviewed. The multiple sequence alignment problem is one the most common task in the analysis of sequential data, especially in bioinformatics. in this paper, we propose to use a genetic algorithm to compute a multiple sequence alignment, by optimizing a simple.

Multiple Sequence Alignment Alchetron The Free Social Encyclopedia In this review, the pairwise sequence alignment algorithms and the corresponding scoring system, heuristic algorithms for multiple sequence alignment and their defects, and quality estimation methods used to test multiple sequence alignment software are reviewed. The multiple sequence alignment problem is one the most common task in the analysis of sequential data, especially in bioinformatics. in this paper, we propose to use a genetic algorithm to compute a multiple sequence alignment, by optimizing a simple. We performed comprehensive searches using keywords related to multiple sequence alignment algorithms across major scientific databases, including pubmed, ieee xplore, and google scholar. A multiple sequence alignment is an alignment of n > 2 sequences obtained by inserting gaps (“‐”) into sequences such that the resulting sequences have all length l and can be arranged in a matrix of n rows and l columns where each column represents a homologous position. Mafft is a multiple sequence alignment program for unix like operating systems. it offers a range of multiple alignment methods, l ins i (accurate; for alignment of <∼200 sequences), fft ns 2 (fast; for alignment of <∼30,000 sequences), etc. Results showed that alignment quality was highly dependent on the number of deletions and insertions in the sequences and that the sequence length and indel size had a weaker effect. overall, probcons was consistently on the top of list of the evaluated msa tools.
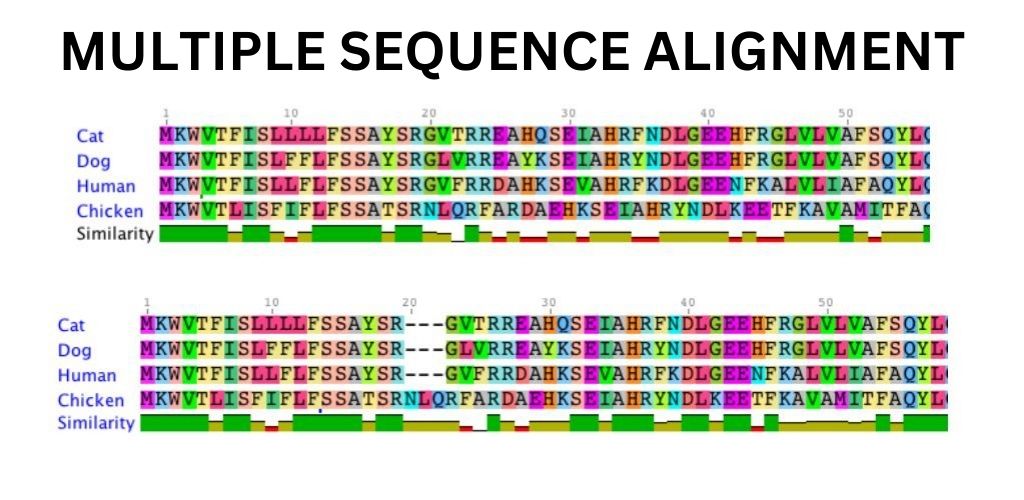
Multiple Sequence Alignment Tools Dna At Tammy Moran Blog We performed comprehensive searches using keywords related to multiple sequence alignment algorithms across major scientific databases, including pubmed, ieee xplore, and google scholar. A multiple sequence alignment is an alignment of n > 2 sequences obtained by inserting gaps (“‐”) into sequences such that the resulting sequences have all length l and can be arranged in a matrix of n rows and l columns where each column represents a homologous position. Mafft is a multiple sequence alignment program for unix like operating systems. it offers a range of multiple alignment methods, l ins i (accurate; for alignment of <∼200 sequences), fft ns 2 (fast; for alignment of <∼30,000 sequences), etc. Results showed that alignment quality was highly dependent on the number of deletions and insertions in the sequences and that the sequence length and indel size had a weaker effect. overall, probcons was consistently on the top of list of the evaluated msa tools.
Github Colmar Zlicheng Multiple Sequence Alignment Dp A Genetic Mafft is a multiple sequence alignment program for unix like operating systems. it offers a range of multiple alignment methods, l ins i (accurate; for alignment of <∼200 sequences), fft ns 2 (fast; for alignment of <∼30,000 sequences), etc. Results showed that alignment quality was highly dependent on the number of deletions and insertions in the sequences and that the sequence length and indel size had a weaker effect. overall, probcons was consistently on the top of list of the evaluated msa tools.
Comments are closed.