Pathogen Agnostic Surveillance With Nanopore Sequencing

Pathogen Agnostic Surveillance With Nanopore Sequencing Biotechniques To address this, we optimized and combined sequence independent single primer amplification (sispa) with depletion of abundant sequences by hybridization (dash), establishing a low cost metagenomic protocol on the oxford nanopore sequencing platform. Building databases to support clinical and research applications of long read sequencing. many approaches for identifying pathogens, such as pcr, typically rely on looking for a specific region by assaying small, highly conserved genomic regions.

Pathogen Agnostic Surveillance With Nanopore Sequencing Biotechniques In this webinar, we explore how oxford nanopore sequencing enables a comprehensive metagenomic approach for characterising bacteria, fungi, and viruses directly from respiratory samples. Conventional pathogen detection methods take 3–7 days and lack sensitivity for low abundance pathogens and antimicrobial resistance (amr) genes. this review evaluates nanopore sequencing's potential in food microbiology, analyzing real time pathogen detection, amr surveillance, and performance versus traditional methods. This review examines the principles of nanopore sequencing and its advantages over conventional methods, surveying recent applications across viral, bacterial (including antimicrobial resistance, amr), and parasitic pathogen detection in animals. This review integrates current nanopore sequencing strategies and applications for animal pathogen detection—across viral, bacterial, and parasitic systems—and places them in the context of outbreak response, surveillance, and genome informed decision making (figure 1).

Accelerating Pathogen Detection With Nanopore Sequencing The Scientist This review examines the principles of nanopore sequencing and its advantages over conventional methods, surveying recent applications across viral, bacterial (including antimicrobial resistance, amr), and parasitic pathogen detection in animals. This review integrates current nanopore sequencing strategies and applications for animal pathogen detection—across viral, bacterial, and parasitic systems—and places them in the context of outbreak response, surveillance, and genome informed decision making (figure 1). Comprehensive pathogen identification, antibiotic resistance, and virulence genes prediction directly from simulated blood samples and positive blood cultures by nanopore metagenomic sequencing. The dhs’s biowatch program demonstrated biothreat agent detection in air samples, and with advanced agnostic sequencing, spectral methods, and ai ml capabilities, can now encompass an expanded pathogen range. Our findings demonstrate that standalone ont sequencing of native dna is sufficiently accurate for routine foodborne pathogen surveillance with current chemistry, software, and appropriate quality control, supporting its use in harmonised genomic surveillance frameworks. A range of nanopore sequencing devices are available to meet the coverage and throughput requirements for all pathogen sequencing applications — from in field sequencing in remote environments to high throughput screening of outbreak samples.
Genomic Pathogen Surveillance With Oxford Nanopore Sequencing The Comprehensive pathogen identification, antibiotic resistance, and virulence genes prediction directly from simulated blood samples and positive blood cultures by nanopore metagenomic sequencing. The dhs’s biowatch program demonstrated biothreat agent detection in air samples, and with advanced agnostic sequencing, spectral methods, and ai ml capabilities, can now encompass an expanded pathogen range. Our findings demonstrate that standalone ont sequencing of native dna is sufficiently accurate for routine foodborne pathogen surveillance with current chemistry, software, and appropriate quality control, supporting its use in harmonised genomic surveillance frameworks. A range of nanopore sequencing devices are available to meet the coverage and throughput requirements for all pathogen sequencing applications — from in field sequencing in remote environments to high throughput screening of outbreak samples.

Genomic Pathogen Surveillance With Oxford Nanopore Sequencing The Our findings demonstrate that standalone ont sequencing of native dna is sufficiently accurate for routine foodborne pathogen surveillance with current chemistry, software, and appropriate quality control, supporting its use in harmonised genomic surveillance frameworks. A range of nanopore sequencing devices are available to meet the coverage and throughput requirements for all pathogen sequencing applications — from in field sequencing in remote environments to high throughput screening of outbreak samples.
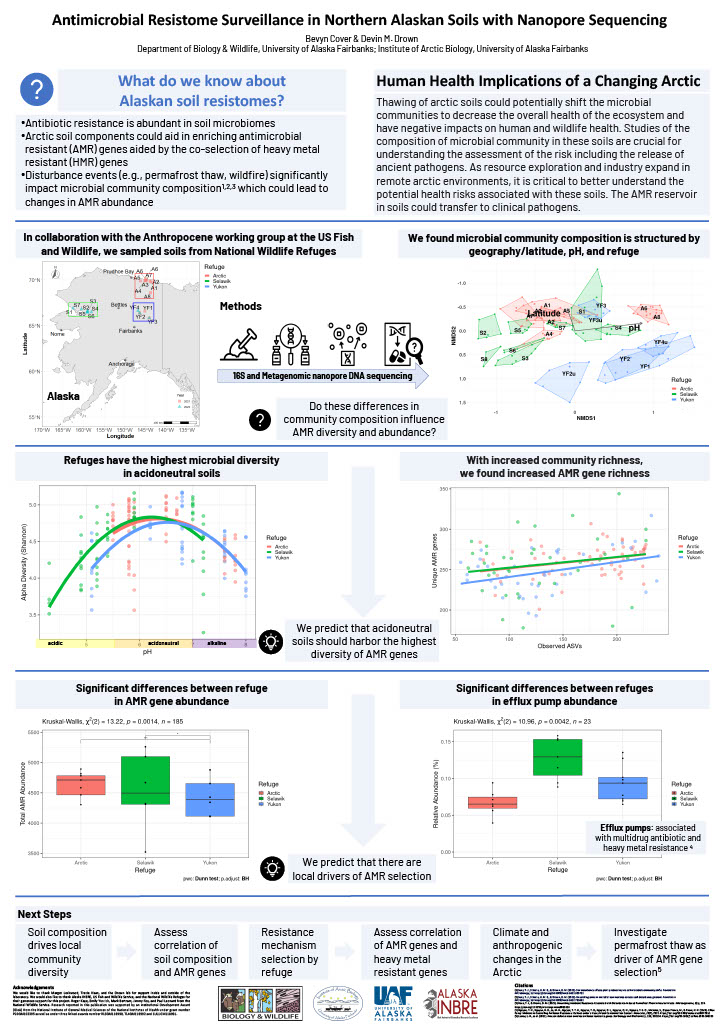
Pathogen Surveillance In Northern Alaskan Soils With Nanopore Sequencing
Comments are closed.