Multivariate Methods For Omics Data Integration And Analysis

Workshop 5 Multivariate Analysis Of Omics Data In this review, we collected the tools and methods that adopt integrative approach to analyze multiple omics data and summarized their ability to address applications such as disease subtyping, biomarker prediction, and deriving insights into the data. In this work, we discuss the application of multivariate statistical approaches to integrate bio molecular information by using multiple factorial analysis. the techniques are applied to a real unpublished data set consisting of three different data types: clinical variables, expression microarrays and dna gel electrophoretic bands.
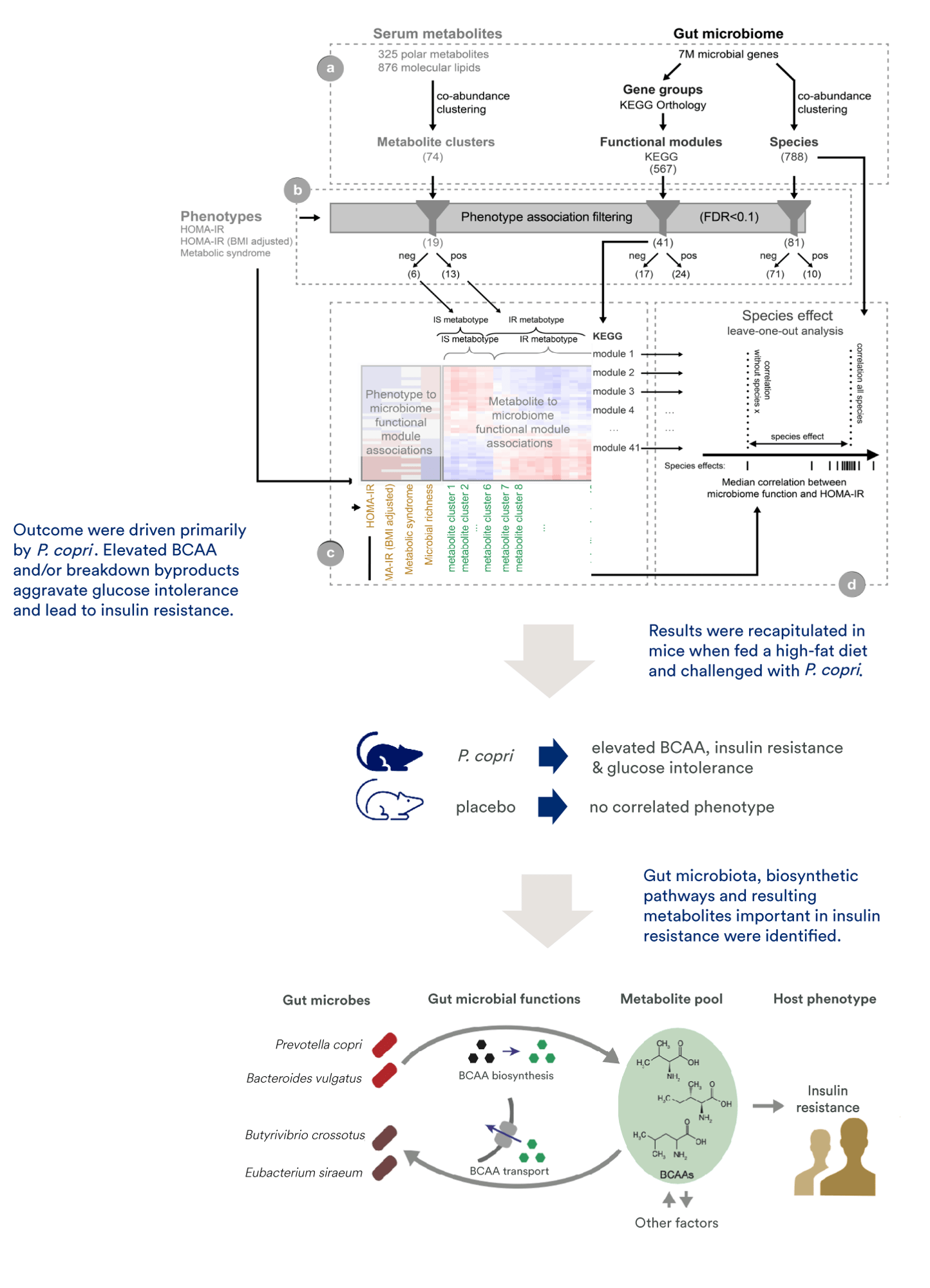
Case Study Multi Omics Data Integration And Insights The mixomics package includes tools for data integration, biomarker discovery, and data visualisation, using advanced multivariate methods to reduce data dimensionality and uncover relationships within and across datasets. Comprehensive review: the paper provides a detailed overview of the multi omics analysis pipeline, covering databases, dimensionality reduction, integration techniques, evaluation metrics, and interpretability and suggests potential improvements and challenges in the field. It starts with some biological background, key concepts underlying the multivariate methods, and then covers an array of methods implemented using the mixomics package in r. In this review, we collected the tools and methods that adopt integrative approach to analyze multiple omics data and summarized their ability to address applications such as disease subtyping, biomarker prediction, and deriving insights into the data.

The Methods For Multi Omics Data Integration Here Simply Shows The It starts with some biological background, key concepts underlying the multivariate methods, and then covers an array of methods implemented using the mixomics package in r. In this review, we collected the tools and methods that adopt integrative approach to analyze multiple omics data and summarized their ability to address applications such as disease subtyping, biomarker prediction, and deriving insights into the data. Here we report the development of the activepathways method that uses data fusion techniques to address the challenge of integrative pathway analysis of multi omics data. We have developed pathintegrate, a pathway based multi omics integration tool which helps integrate and interpret multi omics data in a single step using machine learning. Here, we comprehensively review state of the art multi omics integration methods with a focus on deep generative models, particularly variational autoencoders (vaes) that have been widely used for data imputation, augmentation, and batch effect correction. New biological insights not revealed by single omics analysis (e.g. prostaglandin endoperoxide synthase 2) and pathways common to all data types (interferon, neutrophil degranulation pathways, complement cascade).

The Methods For Multi Omics Data Integration Here Simply Shows The Here we report the development of the activepathways method that uses data fusion techniques to address the challenge of integrative pathway analysis of multi omics data. We have developed pathintegrate, a pathway based multi omics integration tool which helps integrate and interpret multi omics data in a single step using machine learning. Here, we comprehensively review state of the art multi omics integration methods with a focus on deep generative models, particularly variational autoencoders (vaes) that have been widely used for data imputation, augmentation, and batch effect correction. New biological insights not revealed by single omics analysis (e.g. prostaglandin endoperoxide synthase 2) and pathways common to all data types (interferon, neutrophil degranulation pathways, complement cascade).

The Methods For Multi Omics Data Integration Here Simply Shows The Here, we comprehensively review state of the art multi omics integration methods with a focus on deep generative models, particularly variational autoencoders (vaes) that have been widely used for data imputation, augmentation, and batch effect correction. New biological insights not revealed by single omics analysis (e.g. prostaglandin endoperoxide synthase 2) and pathways common to all data types (interferon, neutrophil degranulation pathways, complement cascade).
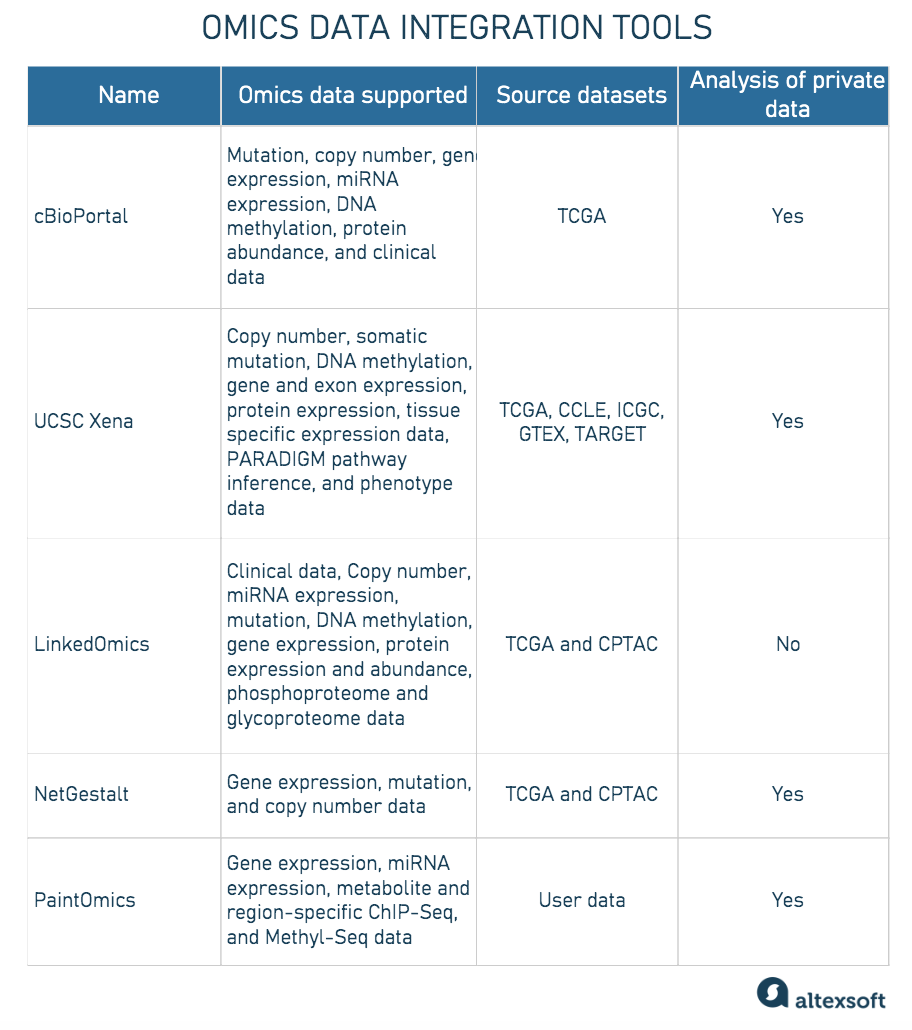
Exploring Omics Data Analysis And Integration Altexsoft
Comments are closed.