Multi Spls Da Comp1 2 Mixomics

Multi Spls Da Comp1 2 Mixomics This page provides a quick start guide for applying partial least squares discriminant analysis (pls da) and its sparse variant (spls da) using mixomics. pls da is the special case of pls where the y dataframe is a single, categorical variable (y). Mixomics is an r toolkit dedicated to the exploration and integration of biological data sets with a specific focus on variable selection. the package currently includes more than twenty multivariate methodologies, mostly developed by the mixomics team (see some of our references in 1.2.3).

Spls Da Comp1 2 Mixomics Mixomics is an r toolkit dedicated to the exploration and integration of biological data sets with a specific focus on variable selection. the package currently includes more than twenty multivariate methodologies, mostly developed by the mixomics team (see some of our references in 1.2.3). Development repository for the bioconductor package 'mixomics ' mixomicsteam mixomics. We present the application of the single ‘omics multivariate methods pca, pls da and spls da on a microarray data set. the pls da and spls da methods are described in the supporting information s1 text. To illustrate the multiblock spls da approach, we will integrate the expression levels of mirna, mrna and the abundance of proteins while discriminating the subtypes of breast cancer, then predict the subtypes of the samples in the test set.
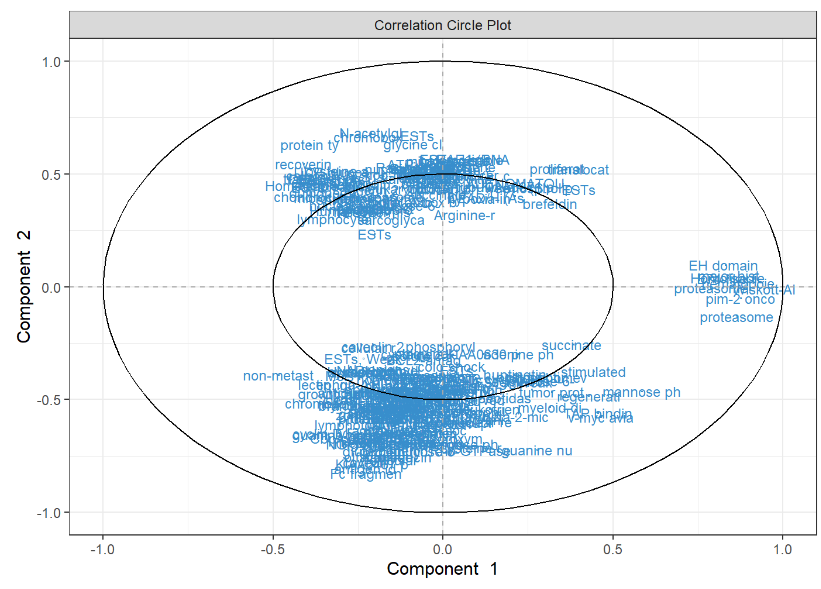
Spls Da Srbct Variable Plot Mixomics We present the application of the single ‘omics multivariate methods pca, pls da and spls da on a microarray data set. the pls da and spls da methods are described in the supporting information s1 text. To illustrate the multiblock spls da approach, we will integrate the expression levels of mirna, mrna and the abundance of proteins while discriminating the subtypes of breast cancer, then predict the subtypes of the samples in the test set. Splsda function fits an spls model with 1, … , ncomp components to the factor or class vector y. the appropriate indicator (dummy) matrix is created. For this engine, there are multiple modes: classification and regression. this model has 2 tuning parameters: the plsmod extension package is required to fit this model. determines the number of predictors in the data. adjusts num comp if the value is larger than the number of factors. Block.splsda function fits a horizontal integration pls da model with a specified number of components per block). a factor indicating the discrete outcome needs to be provided, either by y or by its position indy in the list of blocks x. On spls da: lê cao, k. a., boitard, s. and besse, p. (2011). sparse pls discriminant analysis: biologically relevant feature selection and graphical displays for multiclass problems.
Comments are closed.