Molecule Mapper Devpost

Molecule Mapper Devpost Compound mapper have you ever wanted to map methamphetamine onto the streets of albuquerque? now you can. build real world maps of your favourite compounds to just look at, or recreate on strava!. Researchers at the university of california, santa barbara have developed a genetically encoded protein based sensor called mapper that enables mri to visualize molecular level cellular activity in real time. the system utilizes aquaporin proteins to manipulate water molecule movement, creating a magnetic signal that reveals chemical processes before structural damage occurs. this modular.
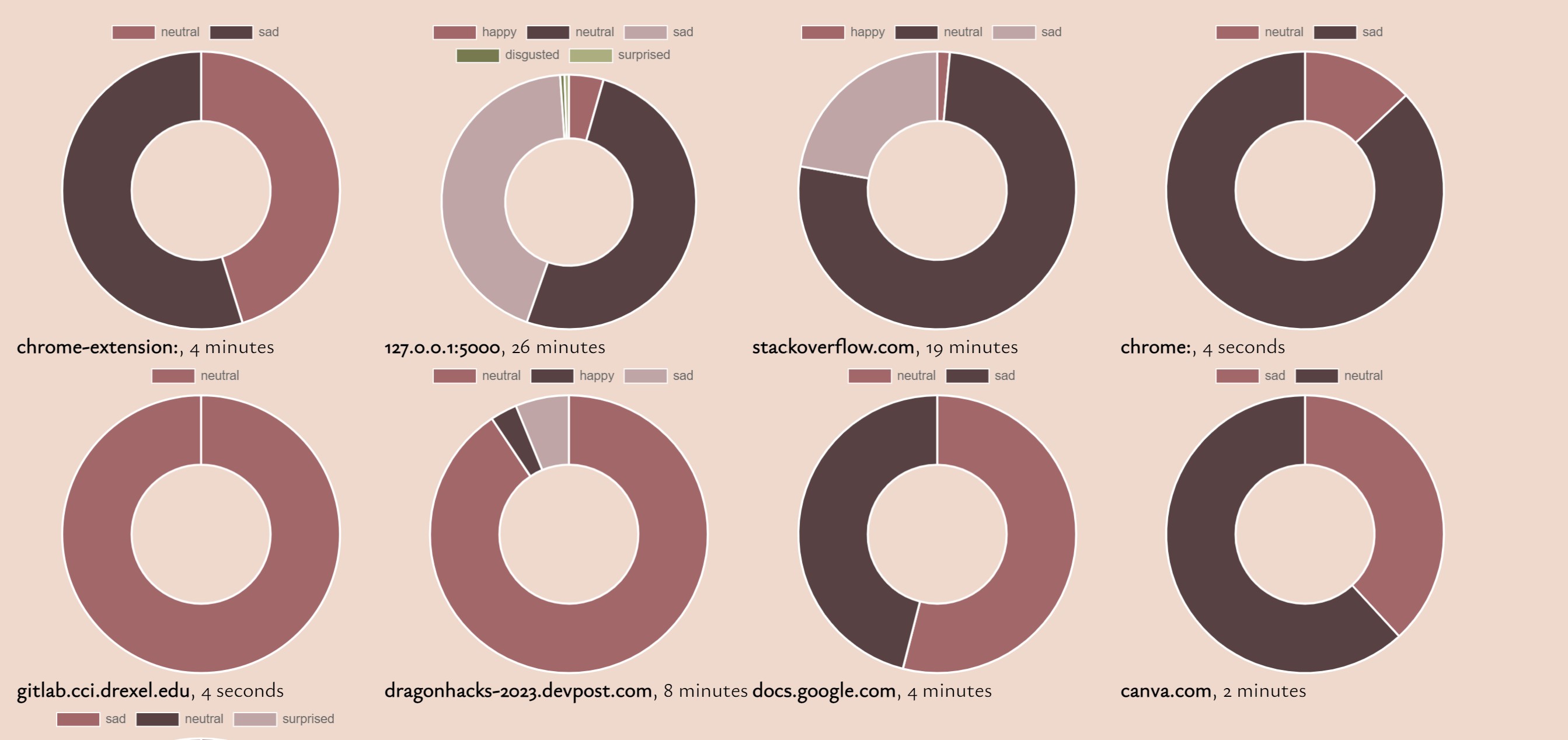
Mood Mapper Devpost Detailed description an accelerated class for rendering molecules. a vtkmoleculemapper that uses imposters to do the rendering. it uses vtkopenglspheremapper and vtkopenglstickmapper to do the rendering. tests: vtkopenglmoleculemapper (tests) definition at line 25 of file vtkopenglmoleculemapper.h. This repository adapts mapper for cheminformatics workflows, integrating molecular structures, scaffolds, physicochemical properties, and geometry based statistics into an interactive web interface. Upon elapsing time limit the molecule is mapped with the so far calculated mapping. parameters: timelimitmilliseconds timeout limit in milliseconds. 1 means disabled. Identifies, calculates and counts the functional groups present in the given rdkit molecule.
Mood Mapper Devpost Upon elapsing time limit the molecule is mapped with the so far calculated mapping. parameters: timelimitmilliseconds timeout limit in milliseconds. 1 means disabled. Identifies, calculates and counts the functional groups present in the given rdkit molecule. When annotators disagree, topology explains: mapper, a topological tool for exploring text embedding geometry and ambiguity nisrine rair, alban goupil, valeriu vrabie, emmanuel chochoy. Recent advances in high throughput molecular methods allow for the collection of complementary methylation and gene expression data from the same sample in large numbers. When multiple molecules map to one node, the corresponding molecular data are summarized into a single node summary by calling function specified by node.sum. this mapped node summary data together with the parsed kgml data are then returned for further processing. One method to evaluate the relationship between dna methylation and gene expression is the mapping of expression quantitative trait methylation (eqtm) loci (also called expression associated cpg loci, ecpg). however, no open source tools are available to provide eqtm mapping.
Comments are closed.